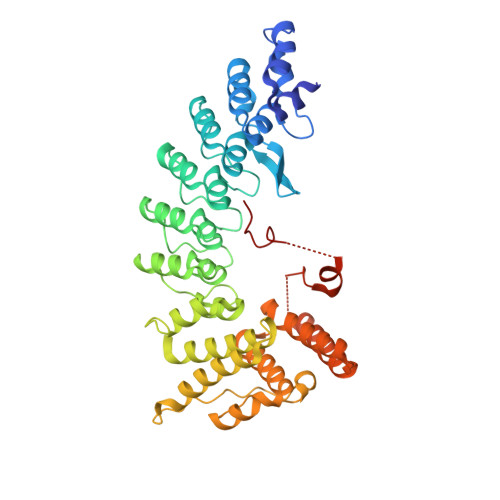
Molecular basis for arginine C-terminal degron recognition by Cul2 FEM1 E3 ligase.
Chen, X., Liao, S., Makaros, Y., Guo, Q., Zhu, Z., Krizelman, R., Dahan, K., Tu, X., Yao, X., Koren, I., Xu, C.(2021) Nat Chem Biol 17: 254-262
- PubMed: 33398168
- DOI: https://doi.org/10.1038/s41589-020-00704-3
- Primary Citation of Related Structures:
6LBF, 6LBG, 6LBN, 6LDP, 6LE6, 6LEN, 6LEY, 6LF0, 7CNG - PubMed Abstract:
Degrons are elements within protein substrates that mediate the interaction with specific degradation machineries to control proteolysis. Recently, a few classes of C-terminal degrons (C-degrons) that are recognized by dedicated cullin-RING ligases (CRLs) have been identified. Specifically, CRL2 using the related substrate adapters FEM1A/B/C was found to recognize C degrons ending with arginine (Arg/C-degron). Here, we uncover the molecular mechanism of Arg/C-degron recognition by solving a subset of structures of FEM1 proteins in complex with Arg/C-degron-bearing substrates. Our structural research, complemented by binding assays and global protein stability (GPS) analyses, demonstrates that FEM1A/C and FEM1B selectively target distinct classes of Arg/C-degrons. Overall, our study not only sheds light on the molecular mechanism underlying Arg/C-degron recognition for precise control of substrate turnover, but also provides valuable information for development of chemical probes for selectively regulating proteostasis.
- MOE Key Laboratory for Membraneless Organelles and Cellular Dynamics, Hefei National Laboratory for Physical Sciences at the Microscale, School of Life Sciences, University of Science and Technology of China, Hefei, China.
Organizational Affiliation: