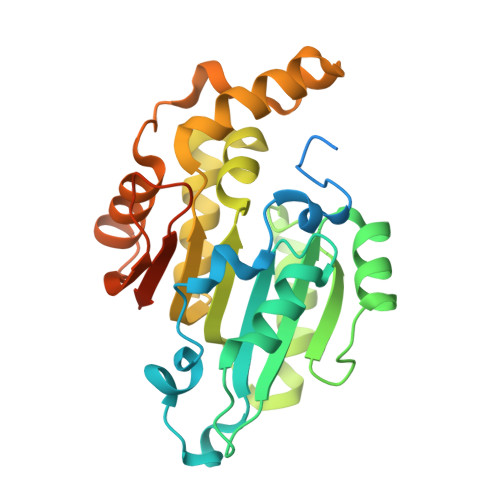
Crystal structure of human cytoplasmic tRNAHis-specific 5'-monomethylphosphate capping enzyme.
Liu, Y., Martinez, A., Yamashita, S., Tomita, K.(2020) Nucleic Acids Res 48: 1572-1582
- PubMed: 31919512 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz1216
- Primary Citation Related Structures:
6L8U - PubMed Abstract:
BCDIN3 domain containing RNA methyltransferase, BCDIN3D, monomethylates the 5'-monophosphate of cytoplasmic tRNAHis with a G-1:A73 mispair at the top of an eight-nucleotide-long acceptor helix, using S-adenosyl-l-methionine (SAM) as a methyl group donor. In humans, BCDIN3D overexpression is associated with the tumorigenic phenotype and poor prognosis in breast cancer. Here, we present the crystal structure of human BCDIN3D complexed with S-adenosyl-l-homocysteine. BCDIN3D adopts a classical Rossmann-fold methyltransferase structure. A comparison of the structure with that of the closely related methylphosphate capping enzyme, MePCE, which monomethylates the 5'-γ-phosphate of 7SK RNA, revealed the important residues for monomethyl transfer from SAM onto the 5'-monophosphate of tRNAHis and for tRNAHis recognition by BCDIN3D. A structural model of tRNAHis docking onto BCDIN3D suggested the molecular mechanism underlying the different activities between BCDIN3D and MePCE. A loop in BCDIN3D is shorter, as compared to the corresponding region that forms an α-helix to recognize the 5'-end of RNA in MePCE, and the G-1:A73 mispair in tRNAHis allows the N-terminal α-helix of BCDIN3D to wedge the G-1:A73 mispair of tRNAHis. As a result, the 5'-monophosphate of G-1 of tRNAHis is deep in the catalytic pocket for 5'-phosphate methylation. Thus, BCDIN3D is a tRNAHis-specific 5'-monomethylphosphate capping enzyme that discriminates tRNAHis from other tRNA species, and the structural information presented in this study also provides the molecular basis for the development of drugs against breast cancers.
- Department of Computational Biology and Medical Sciences, Graduate School of Frontier Sciences, The University of Tokyo, Kashiwa, Chiba 277-8562, Japan.
Organizational Affiliation: