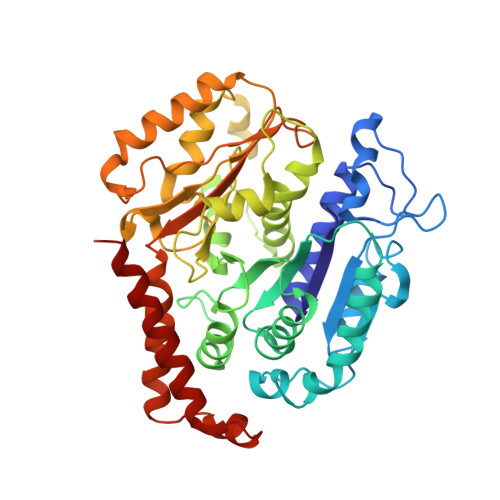
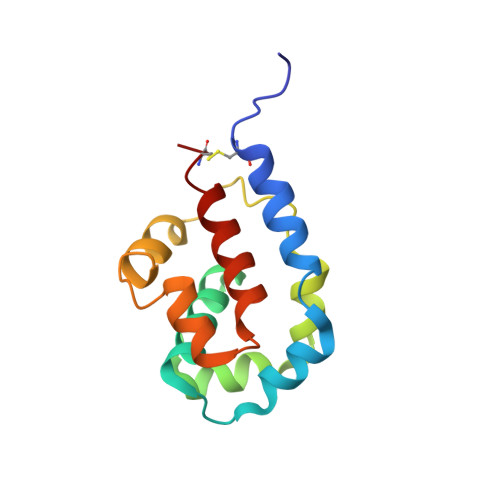
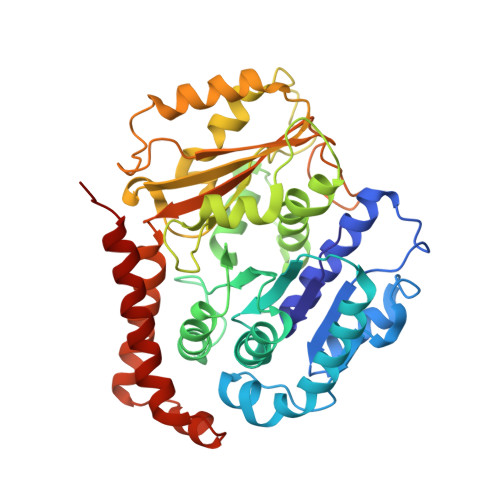
Structural basis for two-way communication between dynein and microtubules.
Nishida, N., Komori, Y., Takarada, O., Watanabe, A., Tamura, S., Kubo, S., Shimada, I., Kikkawa, M.(2020) Nat Commun 11: 1038-1038
- PubMed: 32098965 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-14842-8
- Primary Citation Related Structures:
6KIO, 6KIQ, 6KJN, 6KJO - PubMed Abstract:
The movements of cytoplasmic dynein on microtubule (MT) tracks is achieved by two-way communication between the microtubule-binding domain (MTBD) and the ATPase domain via a coiled-coil stalk, but the structural basis of this communication remains elusive. Here, we regulate MTBD either in high-affinity or low-affinity states by introducing a disulfide bond to the stalk and analyze the resulting structures by NMR and cryo-EM. In the MT-unbound state, the affinity changes of MTBD are achieved by sliding of the stalk α-helix by a half-turn, which suggests that structural changes propagate from the ATPase-domain to MTBD. In addition, MT binding induces further sliding of the stalk α-helix even without the disulfide bond, suggesting how the MT-induced conformational changes propagate toward the ATPase domain. Based on differences in the MT-binding surface between the high- and low-affinity states, we propose a potential mechanism for the directional bias of dynein movement on MT tracks.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo, 113-0033, Japan.
Organizational Affiliation: