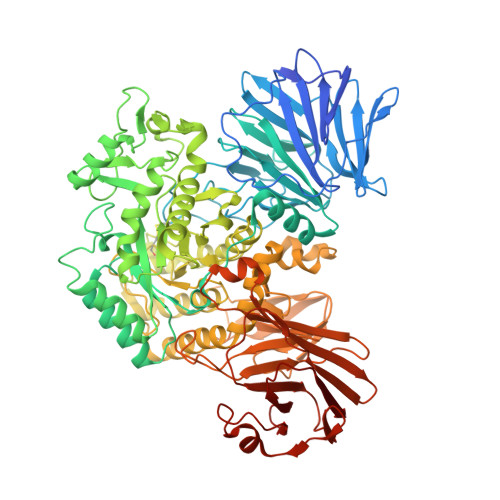
Structural insights into polysaccharide recognition by Flavobacterium johnsoniae dextranase, a member of glycoside hydrolase family 31.
Tsutsumi, K., Gozu, Y., Nishikawa, A., Tonozuka, T.(2020) FEBS J 287: 1195-1207
- PubMed: 31552702 Search on PubMed
- DOI: https://doi.org/10.1111/febs.15074
- Primary Citation Related Structures:
6JR6, 6JR7, 6JR8 - PubMed Abstract:
Glycoside hydrolase family (GH) 31 contains a large variety of enzymes, but the major members are enzymes that act on relatively small oligosaccharides such as α-glucosidase. Here, we determined the crystal structure of Flavobacterium johnsoniae dextranase (FjDex31A), an enzyme from F. johnsoniae that hydrolyzes a polysaccharide, dextran. FjDex31A is composed of four domains: an N-terminal domain, a catalytic domain, a proximal C-terminal domain, and a distal C-terminal domain, as observed in typical GH31 enzymes. However, the architecture of active site residues in FjDex31A, other than subsite -1, is markedly different from that of other GH31 enzymes. The FjDex31A structure in complex with isomaltotriose shows that Gly273 and Tyr524, both of which interact with an α-glucose residue at subsite -2, as well as Trp376 and Leu308-cisGln309, are especially unique to FjDex31A. Site-directed mutagenesis of Gly273 and Tyr524 resulted in a decrease in the hydrolysis of polysaccharides dextran and pullulan, as well as that of the disaccharide isomaltose. These results suggest that, regardless of the length of sugar chains of the substrates, binding of FjDex31A to the substrates at subsite -2 is likely to be important for its activity. DATABASE: Structural data are available in the Protein Data Bank under the accession numbers 6JR6, 6JR7, and 6JR8.
- Department of Applied Biological Science, Tokyo University of Agriculture and Technology, Japan.
Organizational Affiliation: