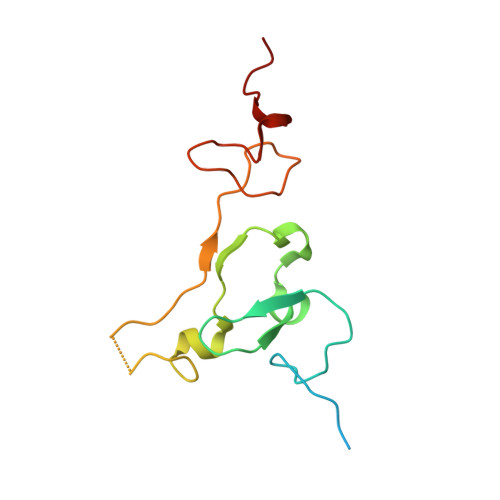
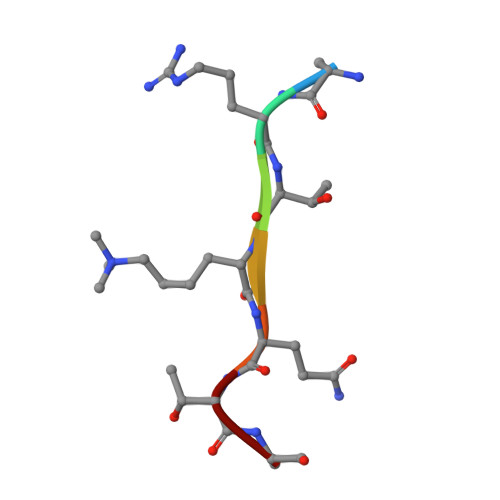
Structural basis for histone H3K4me3 recognition by the N-terminal domain of the PHD finger protein Spp1.
He, C., Liu, N., Xie, D., Liu, Y., Xiao, Y., Li, F.(2019) Biochem J 476: 1957-1973
- PubMed: 31253666 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20190091
- Primary Citation Related Structures:
6J2P - PubMed Abstract:
Saccharomyces cerevisiae Spp1, a plant homeodomain (PHD) finger containing protein, is a critical subunit of the histone H3K4 methyltransferase complex of proteins associated with Set1 (COMPASS). The chromatin binding affinity of the PHD finger of Spp1 has been proposed to modulate COMPASS activity. During meiosis, Spp1 plays another role in promoting programmed double-strand break (DSB) formation by binding H3K4me3 via its PHD finger and interacting with a DSB protein, Mer2. However, how the Spp1 PHD finger performs site-specific readout of H3K4me3 is still not fully understood. In the present study, we determined the crystal structure of the highly conserved Spp1 N-terminal domain (Sc_Spp1 NTD ) in complex with the H3K4me3 peptide. The structure shows that Sc_Spp1 NTD comprises a PHD finger responsible for methylated H3K4 recognition and a C3H-type zinc finger necessary to ensure the overall structural stability. Our isothermal titration calorimetry results show that binding of H3K4me3 to Sc_Spp1 NTD is mildly inhibited by H3R2 methylation, weakened by H3T6 phosphorylation, and abrogated by H3T3 phosphorylation. This histone modification cross-talk, which is conserved in the Saccharomyces pombe and mammalian orthologs of Sc_Spp1 in vitro , can be rationalized structurally and might contribute to the roles of Spp1 in COMPASS activity regulation and meiotic recombination.
- Anhui Key Laboratory of Modern Biomanufacturing and School of Life Sciences, Anhui University, Hefei, Anhui 230601, China chaohe@ahu.edu.cn lifudong@ustc.edu.cn.
Organizational Affiliation: