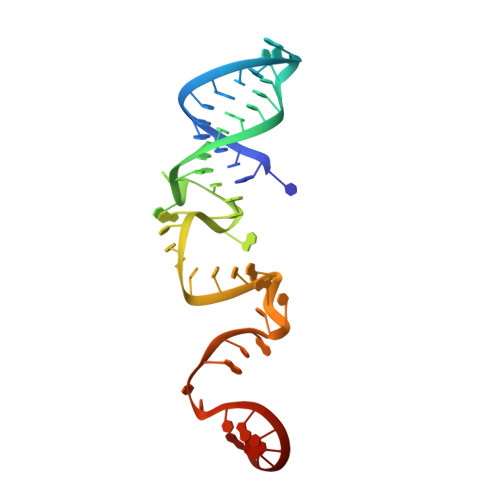
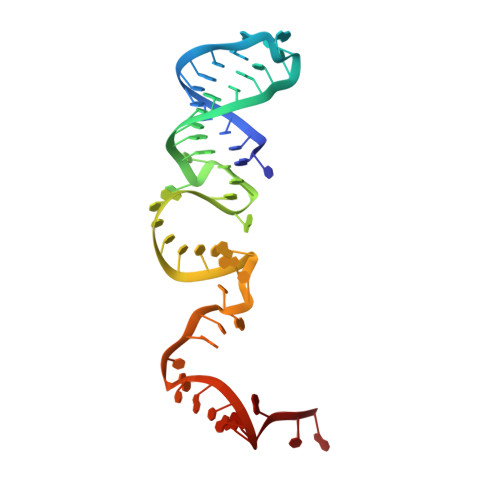
Two HEPN domains dictate CRISPR RNA maturation and target cleavage in Cas13d.
Zhang, B., Ye, Y., Ye, W., Perculija, V., Jiang, H., Chen, Y., Li, Y., Chen, J., Lin, J., Wang, S., Chen, Q., Han, Y.S., Ouyang, S.(2019) Nat Commun 10: 2544-2544
- PubMed: 31186424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-10507-3
- Primary Citation Related Structures:
6IV8, 6IV9 - PubMed Abstract:
Cas13d, the type VI-D CRISPR-Cas effector, is an RNA-guided ribonuclease that has been repurposed to edit RNA in a programmable manner. Here we report the detailed structural and functional analysis of the uncultured Ruminococcus sp. Cas13d (UrCas13d)-crRNA complex. Two hydrated Mg 2+ ions aid in stabilizing the conformation of the crRNA repeat region. Sequestration of divalent metal ions does not alter pre-crRNA processing, but abolishes target cleavage by UrCas13d. Notably, the pre-crRNA processing is executed by the HEPN-2 domain. Furthermore, both the structure and sequence of the nucleotides U(-8)-C(-1) within the repeat region are indispensable for target cleavage, and are specifically recognized by UrCas13d. Moreover, correct base pairings within two separate spacer regions (an internal and a 3'-end region) are essential for target cleavage. These findings provide a framework for the development of Cas13d into a tool for a wide range of applications.
- The Key Laboratory of Innate Immune Biology of Fujian Province, Biomedical Research Center of South China, Key Laboratory of OptoElectronic Science and Technology for Medicine of Ministry of Education, College of Life Sciences, Fujian Normal University, 350117, Fuzhou, China.
Organizational Affiliation: