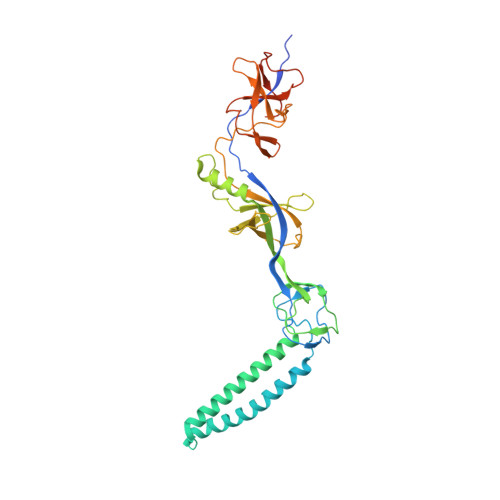
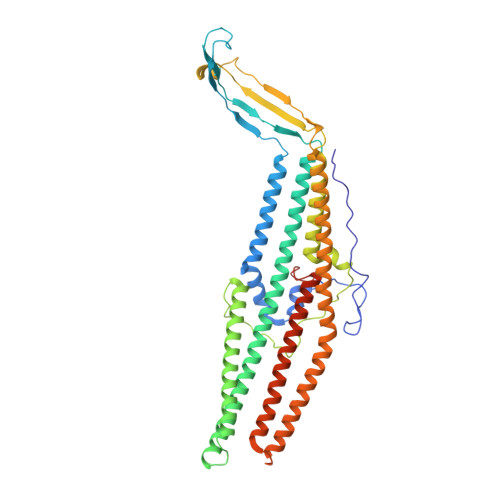
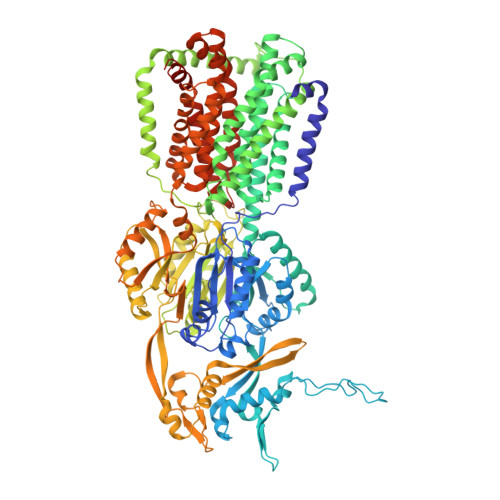
Structures of the wild-type MexAB-OprM tripartite pump reveal its complex formation and drug efflux mechanism.
Tsutsumi, K., Yonehara, R., Ishizaka-Ikeda, E., Miyazaki, N., Maeda, S., Iwasaki, K., Nakagawa, A., Yamashita, E.(2019) Nat Commun 10: 1520-1520
- PubMed: 30944318 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-09463-9
- Primary Citation Related Structures:
6IOK, 6IOL - PubMed Abstract:
In Pseudomonas aeruginosa, MexAB-OprM plays a central role in multidrug resistance by ejecting various drug compounds, which is one of the causes of serious nosocomial infections. Although the structures of the components of MexAB-OprM have been solved individually by X-ray crystallography, no structural information for fully assembled pumps from P. aeruginosa were previously available. In this study, we present the structure of wild-type MexAB-OprM in the presence or absence of drugs at near-atomic resolution. The structure reveals that OprM does not interact with MexB directly, and that it opens its periplasmic gate by forming a complex. Furthermore, we confirm the residues essential for complex formation and observed a movement of the drug entrance gate. Based on these results, we propose mechanisms for complex formation and drug efflux.
- Institute for Protein Research, Osaka University, Suita, 565-0871, Osaka, Japan.
Organizational Affiliation: