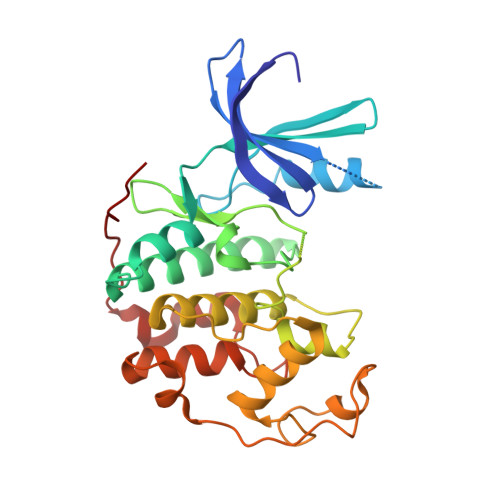
Structure of cyclin-dependent kinase 2 (CDK2) in complex with the specific and potent inhibitor CVT-313.
Talapati, S.R., Nataraj, V., Pothuganti, M., Gore, S., Ramachandra, M., Antony, T., More, S.S., Krishnamurthy, N.R.(2020) Acta Crystallogr F Struct Biol Commun 76: 350-356
- PubMed: 32744246 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X20009243
- Primary Citation Related Structures:
6INL - PubMed Abstract:
CVT-313 is a potent CDK2 inhibitor that was identified by screening a purine-analogue library and is currently in preclinical studies. Since this molecule has the potential to be developed as a CDK2 inhibitor for cancer therapy, the potency of CVT-313 to bind and stabilize CDK2 was evaluated, together with its ability to inhibit aberrant cell proliferation. CVT-313 increased the melting temperature of CDK2 by 7°C in thermal stabilization studies, thus indicating its protein-stabilizing effect. CVT-313 inhibited the growth of human lung carcinoma cell line A549 in a dose-dependent manner, with an IC 50 of 1.2 µM, which is in line with the reported biochemical potency of 0.5 µM. To support the further chemical modification of CVT-313 and to improve its biochemical and cellular potency, a crystal structure was elucidated in order to understand the molecular interaction of CVT-313 and CDK2. The crystal structure of CDK2 bound to CVT-313 was determined to a resolution of 1.74 Å and clearly demonstrated that CVT-313 binds in the ATP-binding pocket, interacting with Leu83, Asp86 and Asp145 directly, and the binding was further stabilized by a water-mediated interaction with Asn132. Based on the crystal structure, further modifications of CVT-313 are proposed to provide additional interactions with CDK2 in the active site, which may significantly increase the biochemical and cellular potency of CVT-313.
- Aurigene Discovery Technologies Ltd, 39-40 KIADB Industrial Area, Electronic City Phase II, Hosur Road, Bangalore 560 100, India.
Organizational Affiliation: