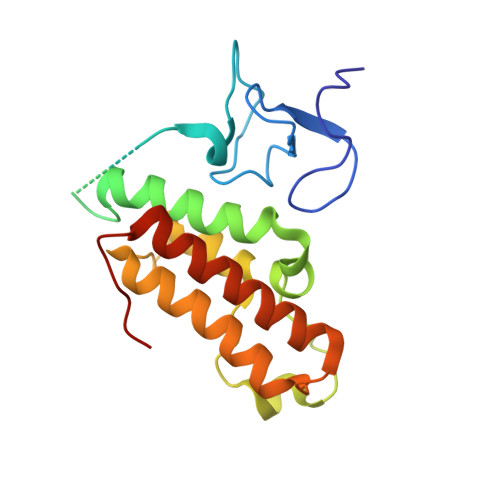
TRIM66 reads unmodified H3R2K4 and H3K56ac to respond to DNA damage in embryonic stem cells.
Chen, J., Wang, Z., Guo, X., Li, F., Wei, Q., Chen, X., Gong, D., Xu, Y., Chen, W., Liu, Y., Kang, J., Shi, Y.(2019) Nat Commun 10: 4273-4273
- PubMed: 31537782 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-12126-4
- Primary Citation Related Structures:
6IET, 6IEU - PubMed Abstract:
Recognition of specific chromatin modifications by distinct structural domains within "reader" proteins plays a critical role in the maintenance of genomic stability. However, the specific mechanisms involved in this process remain unclear. Here we report that the PHD-Bromo tandem domain of tripartite motif-containing 66 (TRIM66) recognizes the unmodified H3R2-H3K4 and acetylated H3K56. The aberrant deletion of Trim66 results in severe DNA damage and genomic instability in embryonic stem cells (ESCs). Moreover, we find that the recognition of histone modification by TRIM66 is critical for DNA damage repair (DDR) in ESCs. TRIM66 recruits Sirt6 to deacetylate H3K56ac, negatively regulating the level of H3K56ac and facilitating the initiation of DDR. Importantly, Trim66-deficient blastocysts also exhibit higher levels of H3K56ac and DNA damage. Collectively, the present findings indicate the vital role of TRIM66 in DDR in ESCs, establishing the relationship between histone readers and maintenance of genomic stability.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui, 230026, China.
Organizational Affiliation: