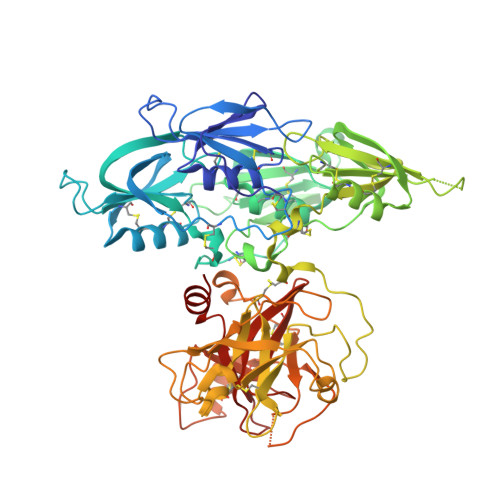
Plasma kallikrein structure reveals apple domain disc rotated conformation compared to factor XI.
Li, C., Voos, K.M., Pathak, M., Hall, G., McCrae, K.R., Dreveny, I., Li, R., Emsley, J.(2019) J Thromb Haemost 17: 759-770
- PubMed: 30801944 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/jth.14418
- Primary Citation Related Structures:
6I44, 6I58, 7QOT, 7QOX - PubMed Abstract:
Essentials Zymogen PK is activated to PKa and cleaves substrates kininogen and FXII contributing to bradykinin generation. Monomeric PKa and dimeric homologue FXI utilize the N-terminal apple domains to recruit substrates. A high-resolution 1.3 Å structure of full-length PKa reveals an active conformation of the protease and apple domains. The PKa protease and four-apple domain disc organization is 180° rotated compared to FXI. SUMMARY: Background Plasma prekallikrein (PK) and factor XI (FXI) are apple domain-containing serine proteases that when activated to PKa and FXIa cleave substrates kininogen, factor XII, and factor IX, respectively, directing plasma coagulation, bradykinin release, inflammation, and thrombosis pathways. Objective To investigate the three-dimensional structure of full-length PKa and perform a comparison with FXI. Methods A series of recombinant full-length PKa and FXI constructs and variants were developed and the crystal structures determined. Results and conclusions A 1.3 Å structure of full-length PKa reveals the protease domain positioned above a disc-shaped assemblage of four apple domains in an active conformation. A comparison with the homologous FXI structure reveals the intramolecular disulfide and structural differences in the apple 4 domain that prevents dimer formation in PK as opposed to FXI. Two latchlike loops (LL1 and LL2) extend from the PKa protease domain to form interactions with the apple 1 and apple 3 domains, respectively. A major unexpected difference in the PKa structure compared to FXI is the 180° disc rotation of the apple domains relative to the protease domain. This results in a switched configuration of the latch loops such that LL2 interacts and buries portions of the apple 3 domain in the FXI zymogen whereas in PKa LL2 interacts with the apple 1 domain. Hydrogen-deuterium exchange mass spectrometry on plasma purified human PK and PKa determined that regions of the apple 3 domain have increased surface exposure in PKa compared to the zymogen PK, suggesting conformational change upon activation.
- Centre for Biomolecular Sciences, School of Pharmacy, University of Nottingham, Nottingham, UK.
Organizational Affiliation: