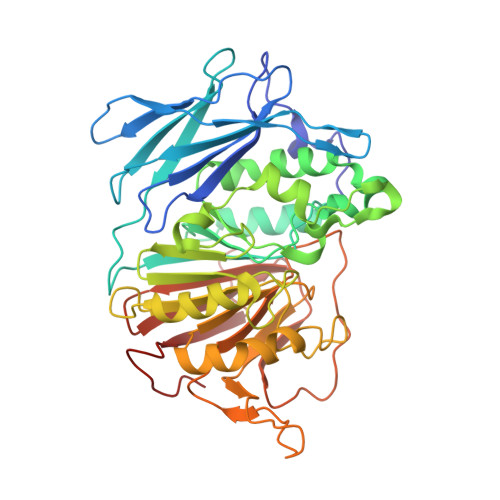
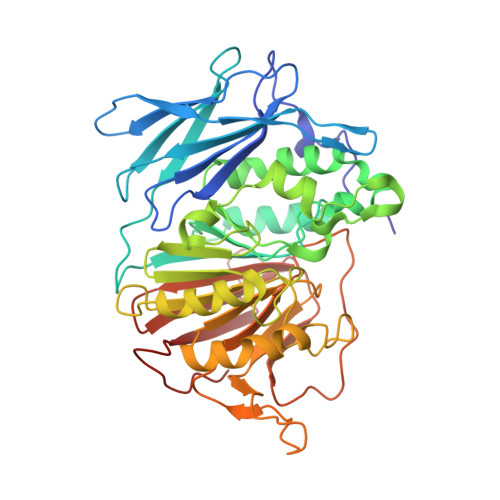
The Binding Mode of an ADP Analogue to a Metallohydrolase Mimics the Likely Transition State.
Feder, D., Gahan, L.R., McGeary, R.P., Guddat, L.W., Schenk, G.(2019) Chembiochem 20: 1536-1540
- PubMed: 30719821 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201900077
- Primary Citation Related Structures:
6HWR - PubMed Abstract:
Purple acid phosphatases (PAPs) are members of the large family of metallohydrolases, a group of enzymes that perform a wide range of biological functions, while employing a highly conserved catalytic mechanism. PAPs are found in plants, animals and fungi; in humans they play an important role in bone turnover and are thus of interest for developing treatments for osteoporosis. The majority of metallohydrolases use a metal-bound hydroxide to initiate catalysis, which leads to the formation of a proposed five-coordinate oxyphosphorane species in the transition state. In this work, we crystallized PAP from red kidney beans (rkbPAP) in the presence of both adenosine and vanadate. The in crystallo-formed vanadate analogue of ADP provides detailed insight into the binding mode of a PAP substrate, captured in a structure that mimics the putative fivecoordinate transition state. Our observations not only provide unprecedented insight into the mechanism of metallohydrolases, but might also guide the structure-based design of inhibitors for application in the treatment of several human illnesses.
- School of Chemistry and Molecular Biosciences, The University of Queensland, St. Lucia, QLD, 4072, Australia.
Organizational Affiliation: