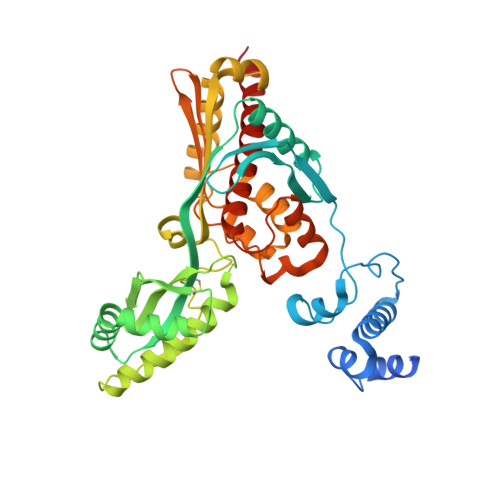
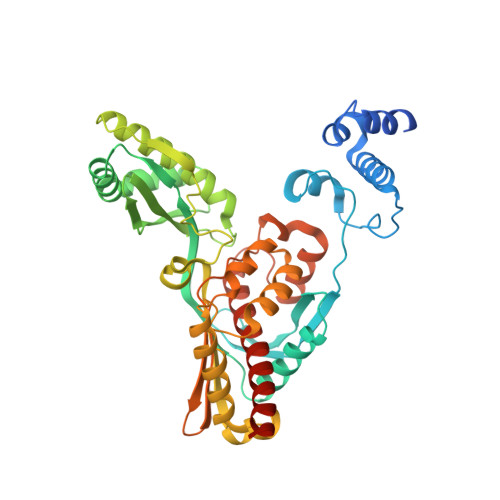
Crystal structure of archaeal HMG-CoA reductase: insights into structural changes of the C-terminal helix of the class-I enzyme.
Vogeli, B., Shima, S., Erb, T.J., Wagner, T.(2019) FEBS Lett 593: 543-553
- PubMed: 30702149 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13331
- Primary Citation Related Structures:
6HR7, 6HR8 - PubMed Abstract:
3-hydroxy-3-methylglutaryl-CoA reductase (HMGR) catalyses the last step in mevalonate biosynthesis. HMGR is the target of statin inhibitors that regulate cholesterol concentration in human blood. Here, we report the properties and structures of HMGR from an archaeon Methanothermococcus thermolithotrophicus (mHMGR). The structures of the apoenzyme and the NADPH complex are highly similar to those of human HMGR. A notable exception is C-terminal helix (Lα10-11) that is straight in both mHMGR structures. This helix is kinked and closes the active site in the human enzyme ternary complex, pointing to a substrate-induced structural rearrangement of C-terminal in class-I HMGRs during the catalytic cycle.
- Department of Biochemistry and Synthetic Metabolism, Max Planck Institute for Terrestrial Microbiology, Marburg, Germany.
Organizational Affiliation: