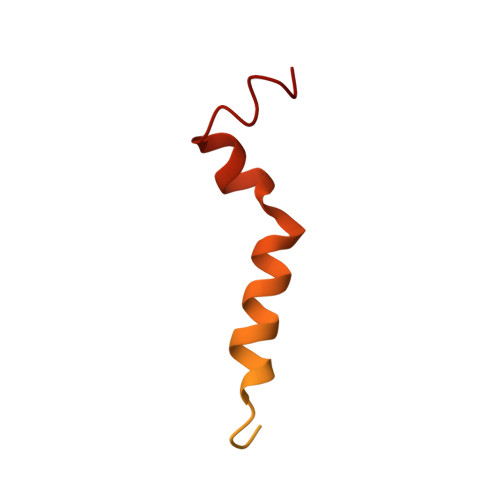
NMR- and MD simulation-based structural characterization of the membrane-associating FATC domain of ataxia telangiectasia mutated.
Abd Rahim, M.S., Cherniavskyi, Y.K., Tieleman, D.P., Dames, S.A.(2019) J Biological Chem 294: 7098-7112
- PubMed: 30867195 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.007653
- Primary Citation Related Structures:
6HKA - PubMed Abstract:
The Ser/Thr protein kinase ataxia telangiectasia mutated (ATM) plays an important role in the DNA damage response, signaling in response to redox signals, the control of metabolic processes, and mitochondrial homeostasis. ATM localizes to the nucleus and at the plasma membrane, mitochondria, peroxisomes, and other cytoplasmic vesicular structures. It has been shown that the C-terminal FATC domain of human ATM (hATMfatc) can interact with a range of membrane mimetics and may thereby act as a membrane-anchoring unit. Here, NMR structural and 15 N relaxation data, NMR data using spin-labeled micelles, and MD simulations of micelle-associated hATMfatc revealed that it binds the micelle by a dynamic assembly of three helices with many residues of hATMfatc located in the headgroup region. We observed that none of the three helices penetrates the micelle deeply or makes significant tertiary contacts to the other helices. NMR-monitored interaction experiments with hATMfatc variants in which two conserved aromatic residues (Phe 3049 and Trp 3052 ) were either individually or both replaced by alanine disclosed that the double substitution does not abrogate the interaction with micelles and bicelles at the high concentrations at which these aggregates are typically used, but impairs interactions with small unilamellar vesicles, usually used at much lower lipid concentrations and considered a better mimetic for natural membranes. We conclude that the observed dynamic structure of micelle-associated hATMfatc may enable it to interact with differently composed membranes or membrane-associated interaction partners and thereby regulate ATM's kinase activity. Moreover, the FATC domain of ATM may function as a membrane-anchoring unit for other biomolecules.
- From the Chair of Biomolecular NMR Spectroscopy, Department of Chemistry, Technische Universität München, Lichtenbergstrasse 4, 85747 Garching, Germany.
Organizational Affiliation: