DNA specificities modulate the binding of human transcription factor A to mitochondrial DNA control region.
Cuppari, A., Fernandez-Millan, P., Battistini, F., Tarres-Sole, A., Lyonnais, S., Iruela, G., Ruiz-Lopez, E., Enciso, Y., Rubio-Cosials, A., Prohens, R., Pons, M., Alfonso, C., Toth, K., Rivas, G., Orozco, M., Sola, M.(2019) Nucleic Acids Res 47: 6519-6537
- PubMed: 31114891 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz406
- Primary Citation Related Structures:
6HB4, 6HC3 - PubMed Abstract:
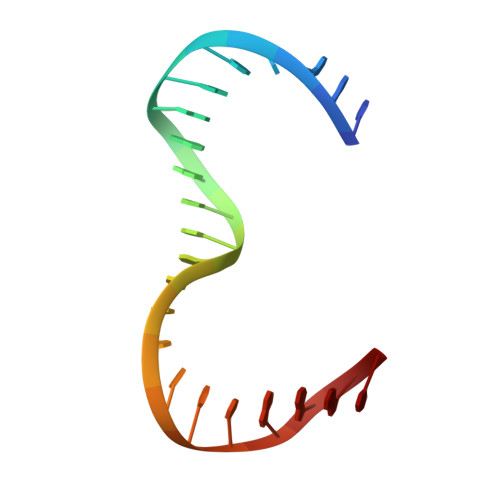
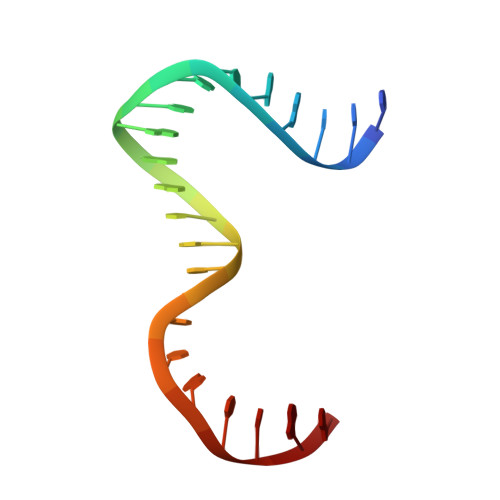
Human mitochondrial DNA (h-mtDNA) codes for 13 subunits of the oxidative phosphorylation pathway, the essential route that produces ATP. H-mtDNA transcription and replication depends on the transcription factor TFAM, which also maintains and compacts this genome. It is well-established that TFAM activates the mtDNA promoters LSP and HSP1 at the mtDNA control region where DNA regulatory elements cluster. Previous studies identified still uncharacterized, additional binding sites at the control region downstream from and slightly similar to LSP, namely sequences X and Y (Site-X and Site-Y) (Fisher et al., Cell 50, pp 247-258, 1987). Here, we explore TFAM binding at these two sites and compare them to LSP by multiple experimental and in silico methods. Our results show that TFAM binding is strongly modulated by the sequence-dependent properties of Site-X, Site-Y and LSP. The high binding versatility of Site-Y or the considerable stiffness of Site-X tune TFAM interactions. In addition, we show that increase in TFAM/DNA complex concentration induces multimerization, which at a very high concentration triggers disruption of preformed complexes. Therefore, our results suggest that mtDNA sequences induce non-uniform TFAM binding and, consequently, direct an uneven distribution of TFAM aggregation sites during the essential process of mtDNA compaction.
- Structural MitoLab, Structural Biology Department, Maria de Maeztu Unit of Excellence, Molecular Biology Institute Barcelona (IBMB-CSIC), 08028 Barcelona, Spain.
Organizational Affiliation: