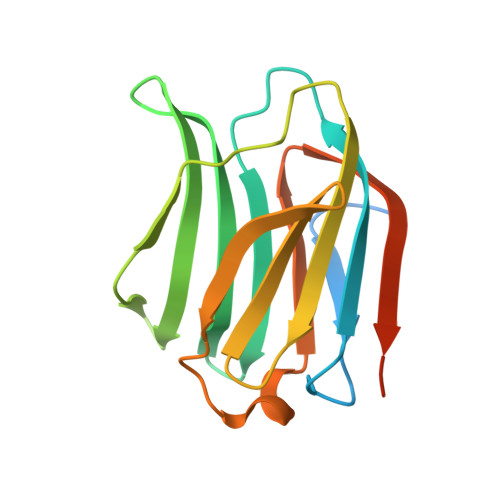
CRD SAT Generated by pCARGHO: A New Efficient Lectin-Based Affinity Tag Method for Safe, Simple, and Low-Cost Protein Purification.
Kriznik, A., Yelehe-Okouma, M., Lec, J.C., Groshenry, G., Le Cordier, H., Charron, C., Quinternet, M., Mazon, H., Talfournier, F., Boschi-Muller, S., Jouzeau, J.Y., Reboul, P.(2019) Biotechnol J 14: e1800214-e1800214
- PubMed: 30298550 Search on PubMed
- DOI: https://doi.org/10.1002/biot.201800214
- Primary Citation Related Structures:
6H64 - PubMed Abstract:
Purification of recombinant proteins remains a bottleneck for downstream processing. The authors engineered a new galectin 3 truncated form (CRD SAT ), functionally and structurally characterized, with preserved solubility and lectinic activity. Taking advantage of these properties, the authors designed an expression vector (pCARGHO), suitable for CRD SAT -tagged protein expression in prokaryotes. CRD SAT binds to lactose-Sepharose with a high specificity and facilitates solubilization of fusion proteins. This tag is structurally stable and can be easily removed from fusion proteins using TEV protease. Furthermore, due to their basic isoelectric point (pI), CRD SAT , and TEV are efficiently eliminated using cationic exchange chromatography. When pI of the protein of interest (POI) and CRD SAT are close, other chromatographic methods are successfully tested. Using CRD SAT tag, the authors purified several proteins from prokaryote and eukaryote origin and demonstrated as examples, the preservation of both Escherichia coli Thioredoxin 1 and human CDC25B cd activities. Overall, yields of proteins obtained after tag removal are about 5-50 mg per litre of bacterial culture. Our purification method displays various advantages described herein that may greatly interest academic laboratories, biotechnology, and pharmaceutical companies.
- Université de Lorraine, CNRS, Ingénierie Moléculaire et Pathologie Articulaire (IMoPA), F-54000, Nancy, France.
Organizational Affiliation: