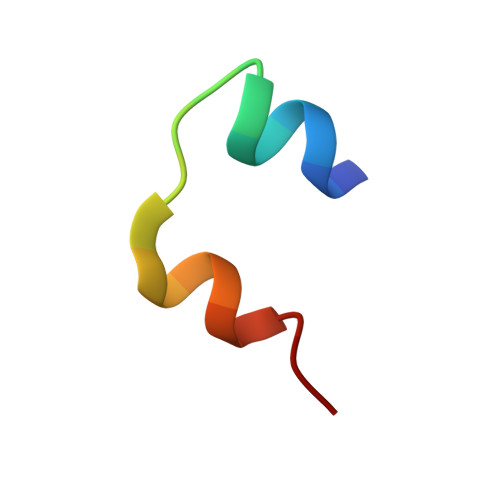
Substitution of an Internal Disulfide Bridge with a Diselenide Enhances both Foldability and Stability of Human Insulin.
Weil-Ktorza, O., Rege, N., Lansky, S., Shalev, D.E., Shoham, G., Weiss, M.A., Metanis, N.(2019) Chemistry 25: 8513-8521
- PubMed: 31012517 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/chem.201900892
- Primary Citation Related Structures:
6H3M - PubMed Abstract:
Insulin analogues, mainstays in the modern treatment of diabetes mellitus, exemplify the utility of protein engineering in molecular pharmacology. Whereas chemical syntheses of the individual A and B chains were accomplished in the early 1960s, their combination to form native insulin remains inefficient because of competing disulfide pairing and aggregation. To overcome these limitations, we envisioned an alternative approach: pairwise substitution of cysteine residues with selenocysteine (Sec, U). To this end, Cys A6 and Cys A11 (which form the internal intrachain A6-A11 disulfide bridge) were each replaced with Sec. The A chain[C6U, C11U] variant was prepared by solid-phase peptide synthesis; while sulfitolysis of biosynthetic human insulin provided wild-type B chain-di-S-sulfonate. The presence of selenium atoms at these sites markedly enhanced the rate and fidelity of chain combination, thus solving a long-standing challenge in chemical insulin synthesis. The affinity of the Se-insulin analogue for the lectin-purified insulin receptor was indistinguishable from that of WT-insulin. Remarkably, the thermodynamic stability of the analogue at 25 °C, as inferred from guanidine denaturation studies, was augmented (ΔΔG u ≈0.8 kcal mol -1 ). In accordance with such enhanced stability, reductive unfolding of the Se-insulin analogue and resistance to enzymatic cleavage by Glu-C protease occurred four times more slowly than that of WT-insulin. 2D-NMR and X-ray crystallographic studies demonstrated a native-like three-dimensional structure in which the diselenide bridge was accommodated in the hydrophobic core without steric clash.
- The Institute of Chemistry, The Hebrew University of Jerusalem, Edmond J. Safra, Givat Ram, Jerusalem, 91904, Israel.
Organizational Affiliation: