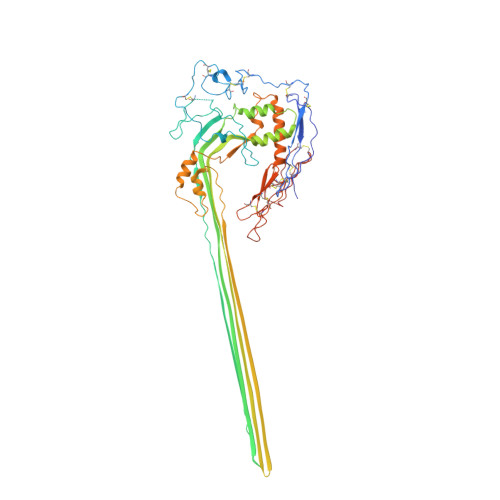
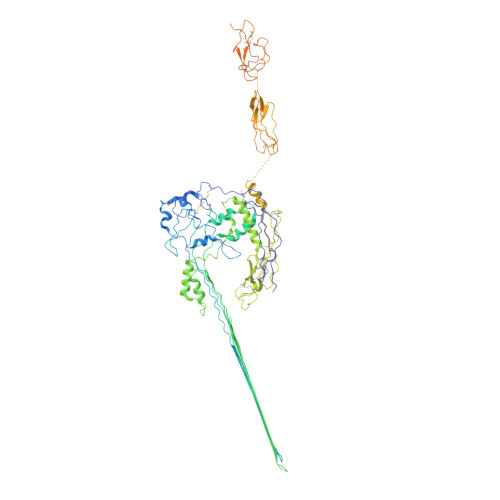
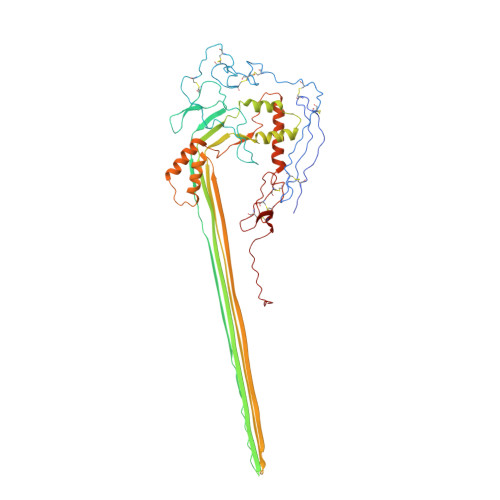
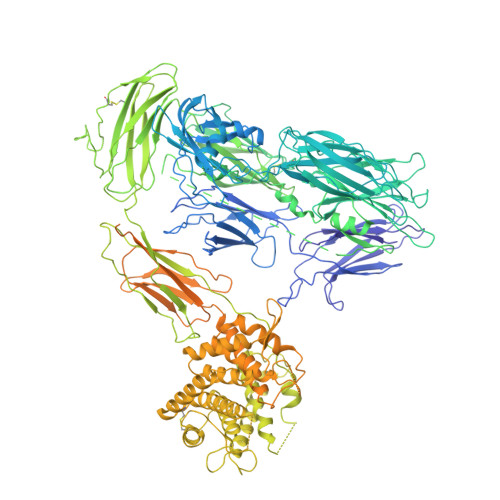
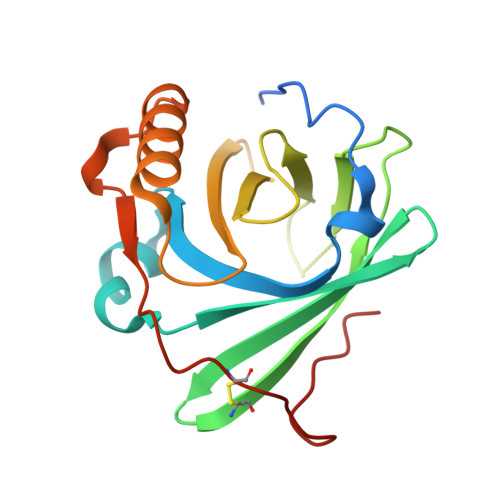
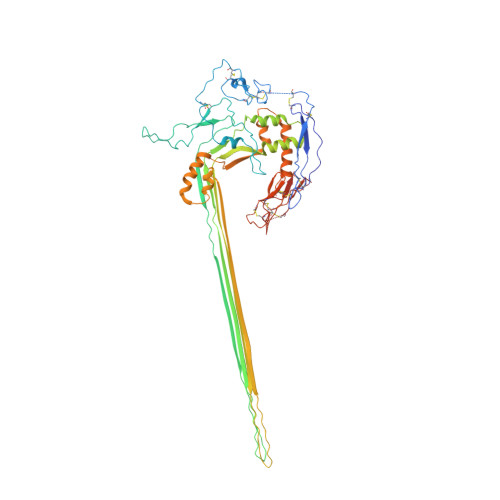
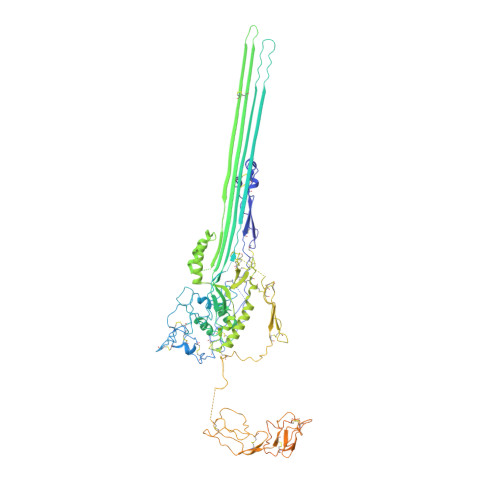
CryoEM reveals how the complement membrane attack complex ruptures lipid bilayers.
Menny, A., Serna, M., Boyd, C.M., Gardner, S., Joseph, A.P., Morgan, B.P., Topf, M., Brooks, N.J., Bubeck, D.(2018) Nat Commun 9: 5316-5316
- PubMed: 30552328 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-07653-5
- Primary Citation Related Structures:
6H03, 6H04 - PubMed Abstract:
The membrane attack complex (MAC) is one of the immune system's first responders. Complement proteins assemble on target membranes to form pores that lyse pathogens and impact tissue homeostasis of self-cells. How MAC disrupts the membrane barrier remains unclear. Here we use electron cryo-microscopy and flicker spectroscopy to show that MAC interacts with lipid bilayers in two distinct ways. Whereas C6 and C7 associate with the outer leaflet and reduce the energy for membrane bending, C8 and C9 traverse the bilayer increasing membrane rigidity. CryoEM reconstructions reveal plasticity of the MAC pore and demonstrate how C5b6 acts as a platform, directing assembly of a giant β-barrel whose structure is supported by a glycan scaffold. Our work provides a structural basis for understanding how β-pore forming proteins breach the membrane and reveals a mechanism for how MAC kills pathogens and regulates cell functions.
- Department of Life Sciences, Sir Ernst Chain Building, Imperial College London, London, SW7 2AZ, UK.
Organizational Affiliation: