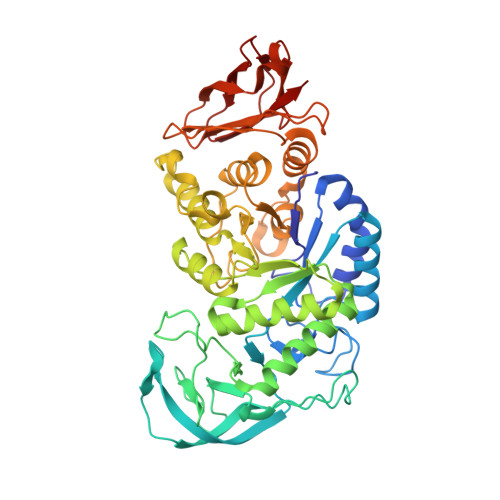
The structure of the AliC GH13 alpha-amylase from Alicyclobacillus sp. reveals the accommodation of starch branching points in the alpha-amylase family.
Agirre, J., Moroz, O., Meier, S., Brask, J., Munch, A., Hoff, T., Andersen, C., Wilson, K.S., Davies, G.J.(2019) Acta Crystallogr D Struct Biol 75: 1-7
- PubMed: 30644839 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798318014900
- Primary Citation Related Structures:
6GXV, 6GYA - PubMed Abstract:
α-Amylases are glycoside hydrolases that break the α-1,4 bonds in starch and related glycans. The degradation of starch is rendered difficult by the presence of varying degrees of α-1,6 branch points and their possible accommodation within the active centre of α-amylase enzymes. Given the myriad industrial uses for starch and thus also for α-amylase-catalysed starch degradation and modification, there is considerable interest in how different α-amylases might accommodate these branches, thus impacting on the potential processing of highly branched post-hydrolysis remnants (known as limit dextrins) and societal applications. Here, it was sought to probe the branch-point accommodation of the Alicyclobacillus sp. CAZy family GH13 α-amylase AliC, prompted by the observation of a molecule of glucose in a position that may represent a branch point in an acarbose complex solved at 2.1 Å resolution. Limit digest analysis by two-dimensional NMR using both pullulan (a regular linear polysaccharide of α-1,4, α-1,4, α-1,6 repeating trisaccharides) and amylopectin starch showed how the Alicyclobacillus sp. enzyme could accept α-1,6 branches in at least the -2, +1 and +2 subsites, consistent with the three-dimensional structures with glucosyl moieties in the +1 and +2 subsites and the solvent-exposure of the -2 subsite 6-hydroxyl group. Together, the work provides a rare insight into branch-point acceptance in these industrial catalysts.
- York Structural Biology Laboratory, Department of Chemistry, The University of York, York YO10 5DD, England.
Organizational Affiliation: