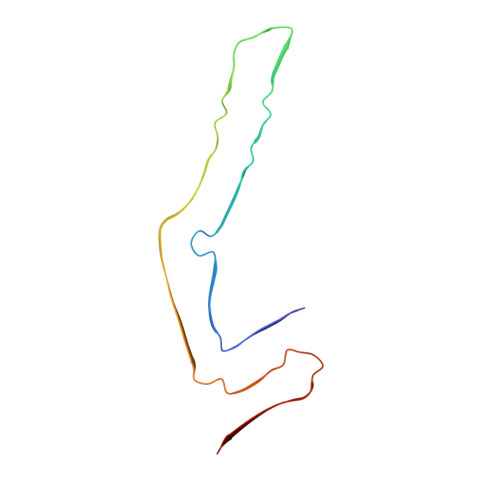
Structures of filaments from Pick's disease reveal a novel tau protein fold.
Falcon, B., Zhang, W., Murzin, A.G., Murshudov, G., Garringer, H.J., Vidal, R., Crowther, R.A., Ghetti, B., Scheres, S.H.W., Goedert, M.(2018) Nature 561: 137-140
- PubMed: 30158706 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0454-y
- Primary Citation Related Structures:
6GX5 - PubMed Abstract:
The ordered assembly of tau protein into abnormal filamentous inclusions underlies many human neurodegenerative diseases 1 . Tau assemblies seem to spread through specific neural networks in each disease 2 , with short filaments having the greatest seeding activity 3 . The abundance of tau inclusions strongly correlates with disease symptoms 4 . Six tau isoforms are expressed in the normal adult human brain-three isoforms with four microtubule-binding repeats each (4R tau) and three isoforms that lack the second repeat (3R tau) 1 . In various diseases, tau filaments can be composed of either 3R or 4R tau, or of both. Tau filaments have distinct cellular and neuroanatomical distributions 5 , with morphological and biochemical differences suggesting that they may be able to adopt disease-specific molecular conformations 6,7 . Such conformers may give rise to different neuropathological phenotypes 8,9 , reminiscent of prion strains 10 . However, the underlying structures are not known. Using electron cryo-microscopy, we recently reported the structures of tau filaments from patients with Alzheimer's disease, which contain both 3R and 4R tau 11 . Here we determine the structures of tau filaments from patients with Pick's disease, a neurodegenerative disorder characterized by frontotemporal dementia. The filaments consist of residues Lys254-Phe378 of 3R tau, which are folded differently from the tau filaments in Alzheimer's disease, establishing the existence of conformers of assembled tau. The observed tau fold in the filaments of patients with Pick's disease explains the selective incorporation of 3R tau in Pick bodies, and the differences in phosphorylation relative to the tau filaments of Alzheimer's disease. Our findings show how tau can adopt distinct folds in the human brain in different diseases, an essential step for understanding the formation and propagation of molecular conformers.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: