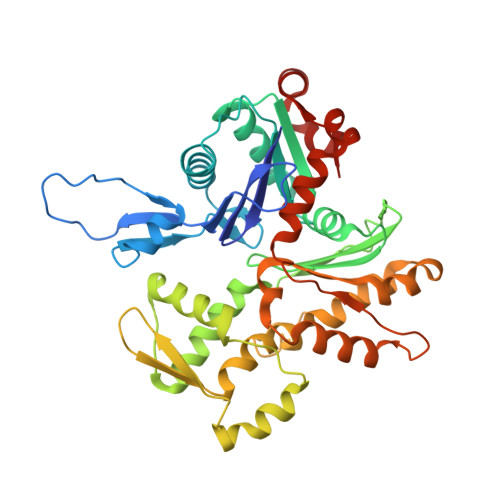
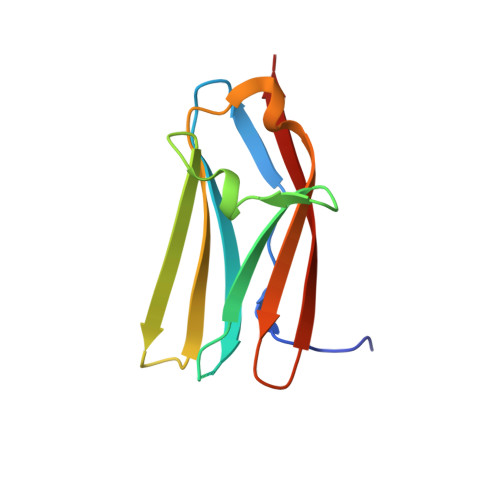
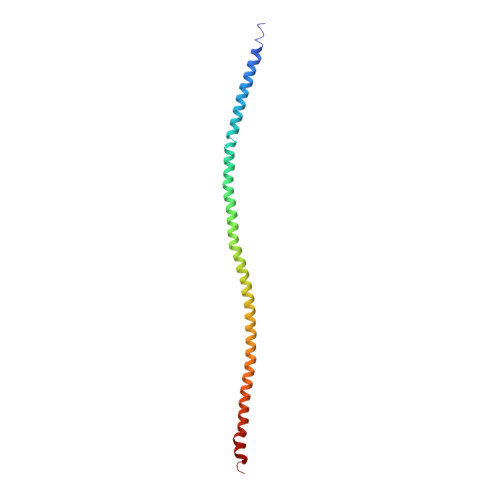N-Terminal Domains of Cardiac Myosin Binding Protein C Cooperatively Activate the Thin Filament.
Risi, C., Belknap, B., Forgacs-Lonart, E., Harris, S.P., Schroder, G.F., White, H.D., Galkin, V.E.(2018) Structure 26: 1604-1611.e4
- PubMed: 30270174 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.08.007
- Primary Citation Related Structures:
6CXI, 6CXJ, 6G2T - PubMed Abstract:
Muscle contraction relies on interaction between myosin-based thick filaments and actin-based thin filaments. Myosin binding protein C (MyBP-C) is a key regulator of actomyosin interactions. Recent studies established that the N'-terminal domains (NTDs) of MyBP-C can either activate or inhibit thin filaments, but the mechanism of their collective action is poorly understood. Cardiac MyBP-C (cMyBP-C) harbors an extra NTD, which is absent in skeletal isoforms of MyBP-C, and its role in regulation of cardiac contraction is unknown. Here we show that the first two domains of human cMyPB-C (i.e., C0 and C1) cooperate to activate the thin filament. We demonstrate that C1 interacts with tropomyosin via a positively charged loop and that this interaction, stabilized by the C0 domain, is required for thin filament activation by cMyBP-C. Our data reveal a mechanism by which cMyBP-C can modulate cardiac contraction and demonstrate a function of the C0 domain.
- Department of Physiological Sciences, Eastern Virginia Medical School, 700 West Olney Road, Lewis Hall, Room 3126, Norfolk, VA 23507, USA.
Organizational Affiliation: