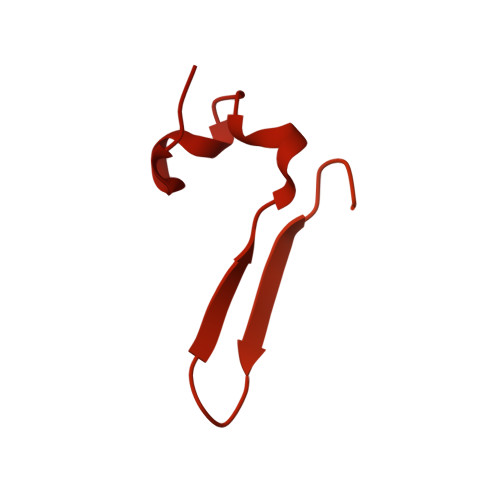
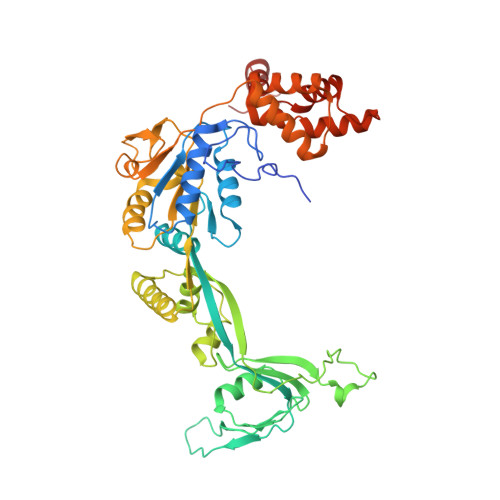
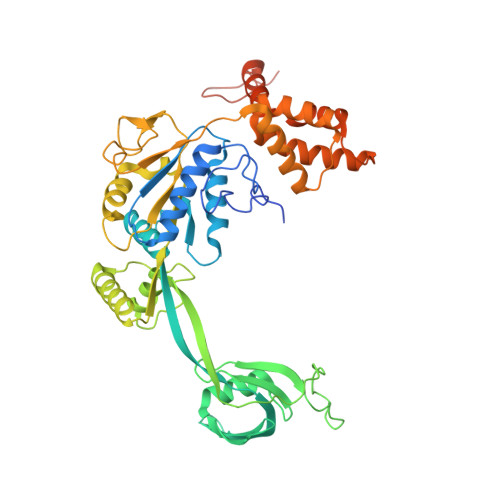
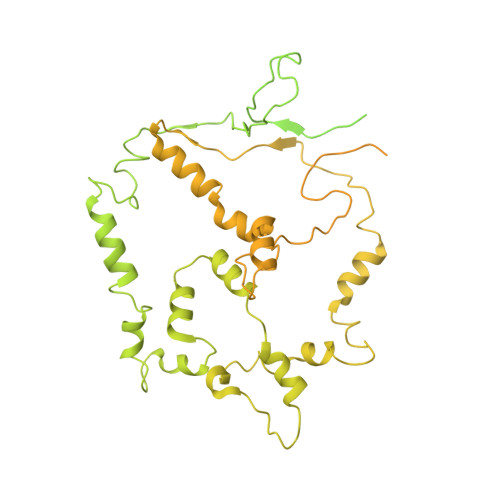
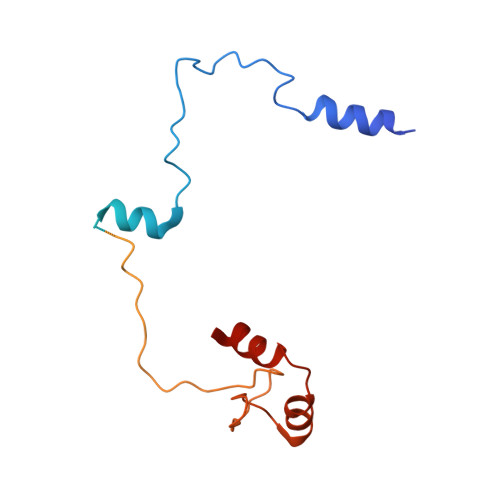
Structural basis for ATP-dependent chromatin remodelling by the INO80 complex.
Eustermann, S., Schall, K., Kostrewa, D., Lakomek, K., Strauss, M., Moldt, M., Hopfner, K.P.(2018) Nature 556: 386-390
- PubMed: 29643509 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-018-0029-y
- Primary Citation Related Structures:
6FHS, 6FML - PubMed Abstract:
In the eukaryotic nucleus, DNA is packaged in the form of nucleosomes, each of which comprises about 147 base pairs of DNA wrapped around a histone protein octamer. The position and histone composition of nucleosomes is governed by ATP-dependent chromatin remodellers 1-3 such as the 15-subunit INO80 complex 4 . INO80 regulates gene expression, DNA repair and replication by sliding nucleosomes, the exchange of histone H2A.Z with H2A, and the positioning of + 1 and -1 nucleosomes at promoter DNA 5-8 . The structures and mechanisms of these remodelling reactions are currently unknown. Here we report the cryo-electron microscopy structure of the evolutionarily conserved core of the INO80 complex from the fungus Chaetomium thermophilum bound to a nucleosome, at a global resolution of 4.3 Å and with major parts at 3.7 Å. The INO80 core cradles one entire gyre of the nucleosome through multivalent DNA and histone contacts. An Rvb1/Rvb2 AAA + ATPase heterohexamer is an assembly scaffold for the complex and acts as a 'stator' for the motor and nucleosome-gripping subunits. The Swi2/Snf2 ATPase motor binds to nucleosomal DNA at superhelical location -6, unwraps approximately 15 base pairs, disrupts the H2A-DNA contacts and is poised to pump entry DNA into the nucleosome. Arp5 and Ies6 bind superhelical locations -2 and -3 to act as a counter grip for the motor, on the other side of the H2A-H2B dimer. The Arp5 insertion domain forms a grappler element that binds the nucleosome dyad, connects the Arp5 actin-fold and entry DNA over a distance of about 90 Å and packs against histone H2A-H2B near the 'acidic patch'. Our structure together with biochemical data 8 suggests a unified mechanism for nucleosome sliding and histone editing by INO80. The motor is part of a macromolecular ratchet, persistently pumping entry DNA across the H2A-H2B dimer against the Arp5 grip until a large nucleosome translocation step occurs. The transient exposure of H2A-H2B by motor activity as well as differential recognition of H2A.Z and H2A may regulate histone exchange.
- Department of Biochemistry, Ludwig-Maximilians-Universität München, Munich, Germany.
Organizational Affiliation: