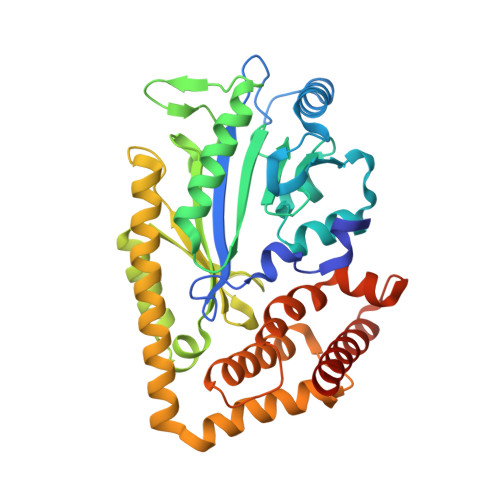
Structure of McsB, a protein kinase for regulated arginine phosphorylation.
Suskiewicz, M.J., Hajdusits, B., Beveridge, R., Heuck, A., Vu, L.D., Kurzbauer, R., Hauer, K., Thoeny, V., Rumpel, K., Mechtler, K., Meinhart, A., Clausen, T.(2019) Nat Chem Biol 15: 510-518
- PubMed: 30962626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-019-0265-y
- Primary Citation Related Structures:
6FH1, 6FH2, 6FH3, 6FH4 - PubMed Abstract:
Protein phosphorylation regulates key processes in all organisms. In Gram-positive bacteria, protein arginine phosphorylation plays a central role in protein quality control by regulating transcription factors and marking aberrant proteins for degradation. Here, we report structural, biochemical, and in vivo data of the responsible kinase, McsB, the founding member of an arginine-specific class of protein kinases. McsB differs in structure and mechanism from protein kinases that act on serine, threonine, and tyrosine residues and instead has a catalytic domain related to that of phosphagen kinases (PhKs), metabolic enzymes that phosphorylate small guanidino compounds. In McsB, the PhK-like phosphotransferase domain is structurally adapted to target protein substrates and is accompanied by a novel phosphoarginine (pArg)-binding domain that allosterically controls protein kinase activity. The identification of distinct pArg reader domains in this study points to a remarkably complex signaling system, thus challenging simplistic views of bacterial protein phosphorylation.
- Research Institute of Molecular Pathology (IMP), Vienna BioCenter (VBC), Vienna, Austria.
Organizational Affiliation: