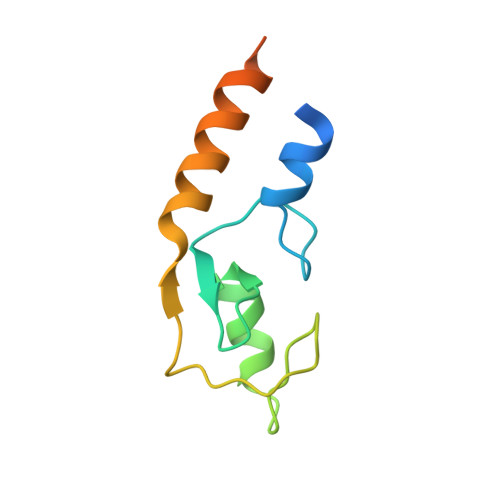
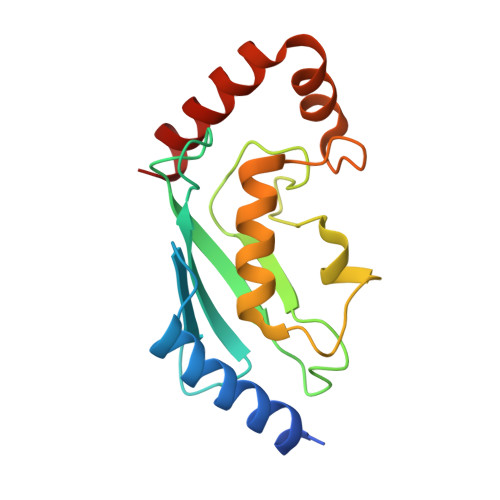
E3 ubiquitin-protein ligase TRIM21-mediated lysine capture by UBE2E1 reveals substrate-targeting mode of a ubiquitin-conjugating E2.
Anandapadamanaban, M., Kyriakidis, N.C., Csizmok, V., Wallenhammar, A., Espinosa, A.C., Ahlner, A., Round, A.R., Trewhella, J., Moche, M., Wahren-Herlenius, M., Sunnerhagen, M.(2019) J Biological Chem 294: 11404-11419
- PubMed: 31160341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.008485
- Primary Citation Related Structures:
6FGA - PubMed Abstract:
The E3 ubiquitin-protein ligase TRIM21, of the RING-containing tripartite motif (TRIM) protein family, is a major autoantigen in autoimmune diseases and a modulator of innate immune signaling. Together with ubiquitin-conjugating enzyme E2 E1 (UBE2E1), TRIM21 acts both as an E3 ligase and as a substrate in autoubiquitination. We here report a 2.82-Å crystal structure of the human TRIM21 RING domain in complex with the human E2-conjugating UBE2E1 enzyme, in which a ubiquitin-targeted TRIM21 substrate lysine was captured in the UBE2E1 active site. The structure revealed that the direction of lysine entry is similar to that described for human proliferating cell nuclear antigen (PCNA), a small ubiquitin-like modifier (SUMO)-targeted substrate, and thus differs from the canonical SUMO-targeted substrate entry. In agreement, we found that critical UBE2E1 residues involved in the capture of the TRIM21 substrate lysine are conserved in ubiquitin-conjugating E2s, whereas residues critical for SUMOylation are not conserved. We noted that coordination of the acceptor lysine leads to remodeling of amino acid side-chain interactions between the UBE2E1 active site and the E2-E3 direct interface, including the so-called "linchpin" residue conserved in RING E3s and required for ubiquitination. The findings of our work support the notion that substrate lysine activation of an E2-E3-connecting allosteric path may trigger catalytic activity and contribute to the understanding of specific lysine targeting by ubiquitin-conjugating E2s.
- Department of Physics, Chemistry and Biology, Division of Chemistry, Linköping University, SE-58183 Linköping, Sweden.
Organizational Affiliation: