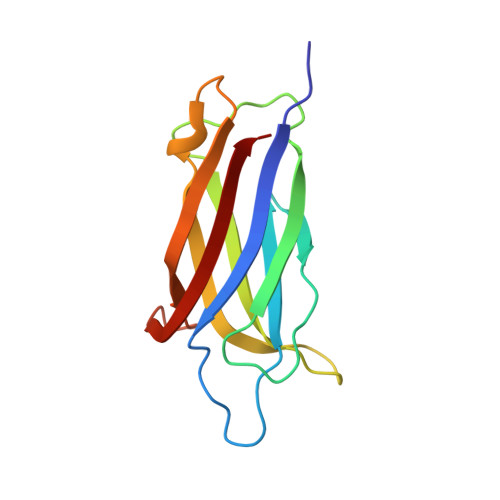
Structural Basis for the Distinct Membrane Binding Activity of the Homologous C2A Domains of Myoferlin and Dysferlin.
Harsini, F.M., Bui, A.A., Rice, A.M., Chebrolu, S., Fuson, K.L., Turtoi, A., Bradberry, M., Chapman, E.R., Sutton, R.B.(2019) J Mol Biology 431: 2112-2126
- PubMed: 31004665 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2019.04.006
- Primary Citation Related Structures:
6EEL - PubMed Abstract:
Dysferlin has been implicated in acute membrane repair processes, whereas myoferlin's activity is maximal during the myoblast fusion stage of early skeletal muscle cell development. Both proteins are similar in size and domain structure; however, despite the overall similarity, myoferlin's known physiological functions do not overlap with those of dysferlin. Here we present for the first time the X-ray crystal structure of human myoferlin C2A to 1.9 Å resolution bound to two divalent cations, and compare its three-dimensional structure and membrane binding activities to that of dysferlin C2A. We find that while dysferlin C2A binds membranes in a Ca 2+ -dependent manner, Ca 2+ binding was the rate-limiting kinetic step for this interaction. Myoferlin C2A, on the other hand, binds two calcium ions with an affinity 3-fold lower than that of dysferlin C2A; and, surprisingly, myoferlin C2A binds only marginally to phospholipid mixtures with a high fraction of phosphatidylserine.
- Department of Cell Physiology and Molecular Biophysics, Texas Tech University Health Sciences Center, Lubbock, TX, 79430, USA; Center for Membrane Protein Research, Texas Tech University Health Sciences Center, Lubbock, TX, 79430, USA.
Organizational Affiliation: