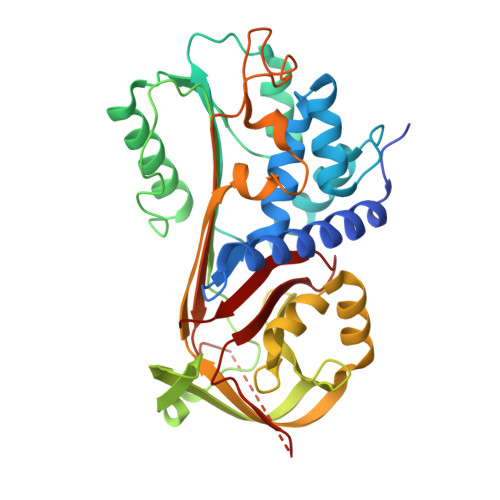
Reactive centre loop dynamics and serpin specificity.
Marijanovic, E.M., Fodor, J., Riley, B.T., Porebski, B.T., Costa, M.G.S., Kass, I., Hoke, D.E., McGowan, S., Buckle, A.M.(2019) Sci Rep 9: 3870-3870
- PubMed: 30846766 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-40432-w
- Primary Citation Related Structures:
6EE5 - PubMed Abstract:
Serine proteinase inhibitors (serpins), typically fold to a metastable native state and undergo a major conformational change in order to inhibit target proteases. However, conformational lability of the native serpin fold renders them susceptible to misfolding and aggregation, and underlies misfolding diseases such as α 1 -antitrypsin deficiency. Serpin specificity towards its protease target is dictated by its flexible and solvent exposed reactive centre loop (RCL), which forms the initial interaction with the target protease during inhibition. Previous studies have attempted to alter the specificity by mutating the RCL to that of a target serpin, but the rules governing specificity are not understood well enough yet to enable specificity to be engineered at will. In this paper, we use conserpin, a synthetic, thermostable serpin, as a model protein with which to investigate the determinants of serpin specificity by engineering its RCL. Replacing the RCL sequence with that from α1-antitrypsin fails to restore specificity against trypsin or human neutrophil elastase. Structural determination of the RCL-engineered conserpin and molecular dynamics simulations indicate that, although the RCL sequence may partially dictate specificity, local electrostatics and RCL dynamics may dictate the rate of insertion during protease inhibition, and thus whether it behaves as an inhibitor or a substrate. Engineering serpin specificity is therefore substantially more complex than solely manipulating the RCL sequence, and will require a more thorough understanding of how conformational dynamics achieves the delicate balance between stability, folding and function required by the exquisite serpin mechanism of action.
- Biomedicine Discovery Institute, Department of Biochemistry and Molecular Biology, Monash University, Victoria, 3800, Australia.
Organizational Affiliation: