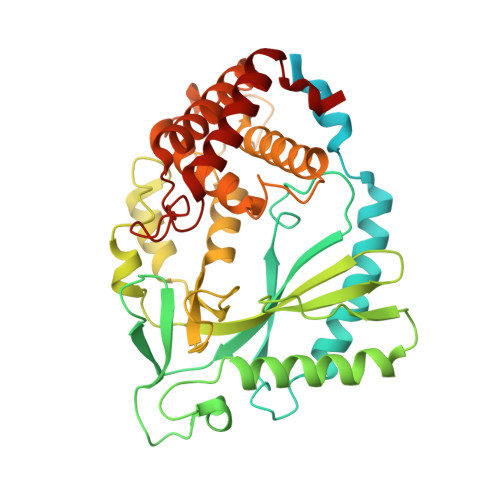
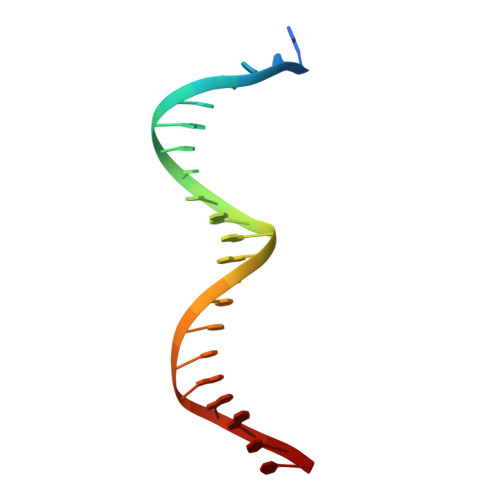
Human cGAS catalytic domain has an additional DNA-binding interface that enhances enzymatic activity and liquid-phase condensation.
Xie, W., Lama, L., Adura, C., Tomita, D., Glickman, J.F., Tuschl, T., Patel, D.J.(2019) Proc Natl Acad Sci U S A 116: 11946-11955
- PubMed: 31142647 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1905013116
- Primary Citation Related Structures:
6EDB, 6EDC, 6O47 - PubMed Abstract:
The cyclic GMP-AMP synthase (cGAS)-cGAMP-STING pathway plays a key role in innate immunity, with cGAS sensing both pathogenic and mislocalized DNA in the cytoplasm. Human cGAS (h-cGAS) constitutes an important drug target for control of antiinflammatory responses that can contribute to the onset of autoimmune diseases. Recent studies have established that the positively charged N-terminal segment of cGAS contributes to enhancement of cGAS enzymatic activity as a result of DNA-induced liquid-phase condensation. We have identified an additional cGAS CD -DNA interface (labeled site-C; CD, catalytic domain) in the crystal structure of a human SRY.cGAS CD -DNA complex, with mutations along this basic site-C cGAS interface disrupting liquid-phase condensation, as monitored by cGAMP formation, gel shift, spin-down, and turbidity assays, as well as time-lapse imaging of liquid droplet formation. We expand on an earlier ladder model of cGAS dimers bound to a pair of parallel-aligned DNAs to propose a multivalent interaction-mediated cluster model to account for DNA-mediated condensation involving both the N-terminal domain of cGAS and the site-C cGAS-DNA interface. We also report the crystal structure of the h-cGAS CD -DNA complex containing a triple mutant that disrupts the site-C interface, with this complex serving as a future platform for guiding cGAS inhibitor development at the DNA-bound h-cGAS level. Finally, we solved the structure of RU.521 bound in two alternate alignments to apo h-cGAS CD , thereby occupying more of the catalytic pocket and providing insights into further optimization of active-site-binding inhibitors.
- Structural Biology Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065; xiew@mskcc.org pateld@mskcc.org.
Organizational Affiliation: