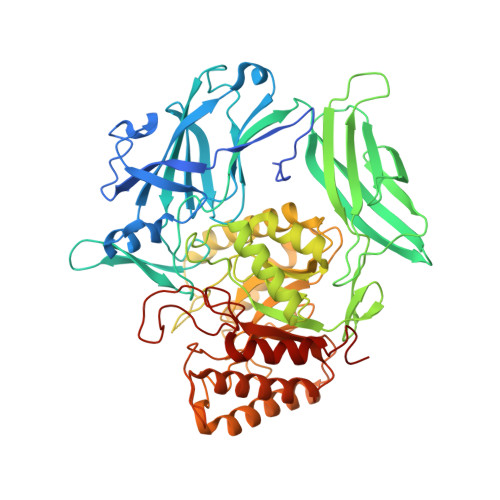
Structure, function, and inhibition of drug reactivating human gut microbial beta-glucuronidases.
Biernat, K.A., Pellock, S.J., Bhatt, A.P., Bivins, M.M., Walton, W.G., Tran, B.N.T., Wei, L., Snider, M.C., Cesmat, A.P., Tripathy, A., Erie, D.A., Redinbo, M.R.(2019) Sci Rep 9: 825-825
- PubMed: 30696850 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-36069-w
- Primary Citation Related Structures:
6EC6, 6ECA, 6ED1, 6ED2 - PubMed Abstract:
Bacterial β-glucuronidase (GUS) enzymes cause drug toxicity by reversing Phase II glucuronidation in the gastrointestinal tract. While many human gut microbial GUS enzymes have been examined with model glucuronide substrates like p-nitrophenol-β-D-glucuronide (pNPG), the GUS orthologs that are most efficient at processing drug-glucuronides remain unclear. Here we present the crystal structures of GUS enzymes from human gut commensals Lactobacillus rhamnosus, Ruminococcus gnavus, and Faecalibacterium prausnitzii that possess an active site loop (Loop 1; L1) analogous to that found in E. coli GUS, which processes drug substrates. We also resolve the structure of the No Loop GUS from Bacteroides dorei. We then compare the pNPG and diclofenac glucuronide processing abilities of a panel of twelve structurally diverse GUS proteins, and find that the new L1 GUS enzymes presented here process small glucuronide substrates inefficiently compared to previously characterized L1 GUS enzymes like E. coli GUS. We further demonstrate that our GUS inhibitors, which are effective against some L1 enzymes, are not potent towards all. Our findings pinpoint active site structural features necessary for the processing of drug-glucuronide substrates and the inhibition of such processing.
- Department of Chemistry, University of North Carolina at Chapel Hill, Chapel Hill, NC, 27599, USA.
Organizational Affiliation: