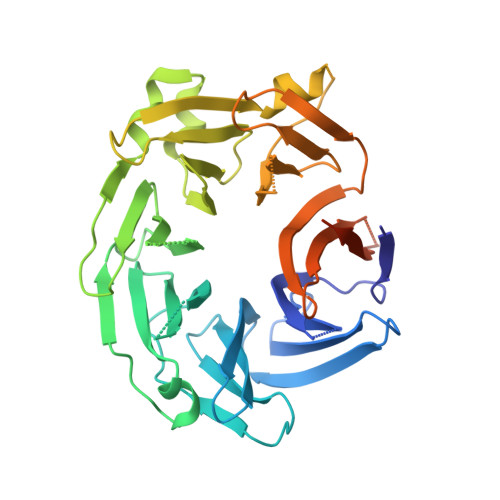
Crystal structure of the WD40 domain dimer of LRRK2.
Zhang, P., Fan, Y., Ru, H., Wang, L., Magupalli, V.G., Taylor, S.S., Alessi, D.R., Wu, H.(2019) Proc Natl Acad Sci U S A 116: 1579-1584
- PubMed: 30635421 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1817889116
- Primary Citation Related Structures:
6DLO, 6DLP - PubMed Abstract:
Leucine-rich repeat kinase 2 (LRRK2) is a large multidomain protein with both a Ras of complex (ROC) domain and a kinase domain (KD) and, therefore, exhibits both GTPase and kinase activities. Human genetics studies have linked LRRK2 as a major genetic contributor to familial and sporadic Parkinson's disease (PD), a neurodegenerative movement disorder that inflicts millions worldwide. The C-terminal region of LRRK2 is a Trp-Asp-40 (WD40) domain with poorly defined biological functions but has been implicated in microtubule interaction. Here, we present the crystal structure of the WD40 domain of human LRRK2 at 2.6-Å resolution, which reveals a seven-bladed WD40 fold. The structure displays a dimeric assembly in the crystal, which we further confirm by measurements in solution. We find that structure-based and PD-associated disease mutations in the WD40 domain including the common G2385R polymorphism mainly compromise dimer formation. Assessment of full-length LRRK2 kinase activity by measuring phosphorylation of Rab10, a member of the family of Rab GTPases known to be important kinase substrates of LRRK2, shows enhancement of kinase activity by several dimerization-defective mutants including G2385R, although dimerization impairment does not always result in kinase activation. Furthermore, mapping of phylogenetically conserved residues onto the WD40 domain structure reveals surface patches that may be important for additional functions of LRRK2. Collectively, our analyses provide insights for understanding the structures and functions of LRRK2 and suggest the potential utility of LRRK2 kinase inhibitors in treating PD patients with WD40 domain mutations.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115.
Organizational Affiliation: