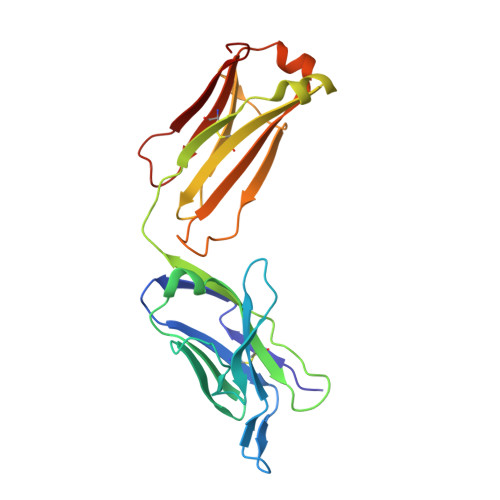
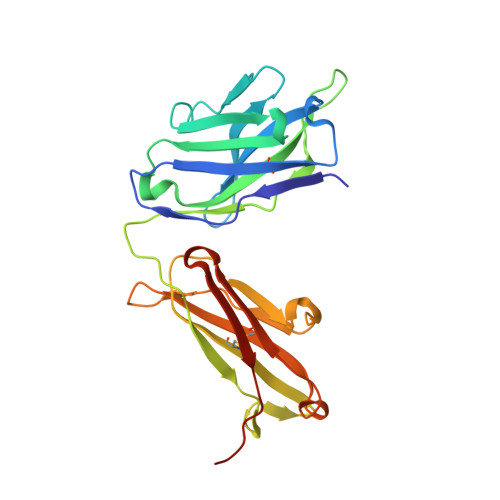
A common antigenic motif recognized by naturally occurring human VH5-51/VL4-1 anti-tau antibodies with distinct functionalities.
Apetri, A., Crespo, R., Juraszek, J., Pascual, G., Janson, R., Zhu, X., Zhang, H., Keogh, E., Holland, T., Wadia, J., Verveen, H., Siregar, B., Mrosek, M., Taggenbrock, R., Ameijde, J., Inganas, H., van Winsen, M., Koldijk, M.H., Zuijdgeest, D., Borgers, M., Dockx, K., Stoop, E.J.M., Yu, W., Brinkman-van der Linden, E.C., Ummenthum, K., van Kolen, K., Mercken, M., Steinbacher, S., de Marco, D., Hoozemans, J.J., Wilson, I.A., Koudstaal, W., Goudsmit, J.(2018) Acta Neuropathol Commun 6: 43-43
- PubMed: 29855358 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s40478-018-0543-z
- Primary Citation Related Structures:
5ZV3, 6DCV, 6DCW, 6GK7, 6GK8 - PubMed Abstract:
Misfolding and aggregation of tau protein are closely associated with the onset and progression of Alzheimer's Disease (AD). By interrogating IgG + memory B cells from asymptomatic donors with tau peptides, we have identified two somatically mutated V H 5-51/V L 4-1 antibodies. One of these, CBTAU-27.1, binds to the aggregation motif in the R3 repeat domain and blocks the aggregation of tau into paired helical filaments (PHFs) by sequestering monomeric tau. The other, CBTAU-28.1, binds to the N-terminal insert region and inhibits the spreading of tau seeds and mediates the uptake of tau aggregates into microglia by binding PHFs. Crystal structures revealed that the combination of V H 5-51 and V L 4-1 recognizes a common Pro-X n -Lys motif driven by germline-encoded hotspot interactions while the specificity and thereby functionality of the antibodies are defined by the CDR3 regions. Affinity improvement led to improvement in functionality, identifying their epitopes as new targets for therapy and prevention of AD.
- Janssen Prevention Center, Janssen Pharmaceutical Companies of Johnson & Johnson, Archimedesweg 6, 2333, CN, Leiden, the Netherlands. AApetri@its.jnj.com.
Organizational Affiliation: