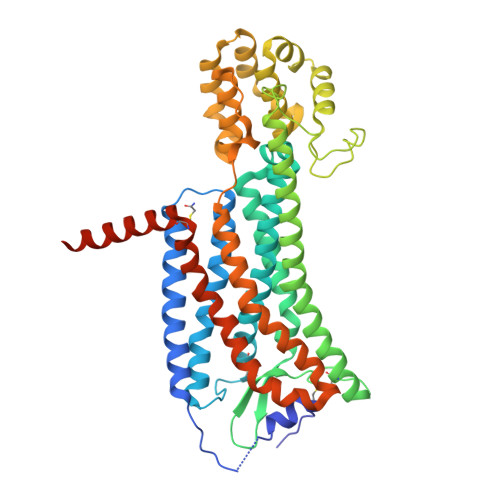
Structures of the Human PGD2Receptor CRTH2 Reveal Novel Mechanisms for Ligand Recognition.
Wang, L., Yao, D., Deepak, R.N.V.K., Liu, H., Xiao, Q., Fan, H., Gong, W., Wei, Z., Zhang, C.(2018) Mol Cell 72: 48-59.e4
- PubMed: 30220562 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2018.08.009
- Primary Citation Related Structures:
6D26, 6D27 - PubMed Abstract:
The signaling of prostaglandin D 2 (PGD 2 ) through G-protein-coupled receptor (GPCR) CRTH2 is a major pathway in type 2 inflammation. Compelling evidence suggests the therapeutic benefits of blocking CRTH2 signaling in many inflammatory disorders. Currently, a number of CRTH2 antagonists are under clinical investigation, and one compound, fevipiprant, has advanced to phase 3 clinical trials for asthma. Here, we present the crystal structures of human CRTH2 with two antagonists, fevipiprant and CAY10471. The structures, together with docking and ligand-binding data, reveal a semi-occluded pocket covered by a well-structured amino terminus and different binding modes of chemically diverse CRTH2 antagonists. Structural analysis suggests a ligand entry port and a binding process that is facilitated by opposite charge attraction for PGD 2 , which differs significantly from the binding pose and binding environment of lysophospholipids and endocannabinoids, revealing a new mechanism for lipid recognition by GPCRs.
- Department of Pharmacology and Chemical Biology, School of Medicine, University of Pittsburgh, Pittsburgh, PA 15261, USA.
Organizational Affiliation: