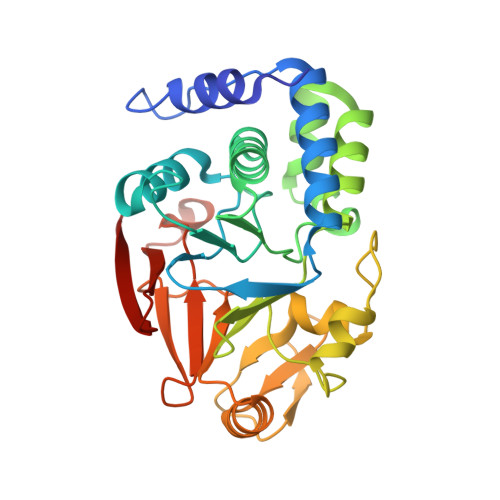
KNL1 Binding to PP1 and Microtubules Is Mutually Exclusive.
Bajaj, R., Bollen, M., Peti, W., Page, R.(2018) Structure 26: 1327-1336.e4
- PubMed: 30100357 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2018.06.013
- Primary Citation Related Structures:
6CZO - PubMed Abstract:
The kinetochore scaffold 1 (KNL1) protein coordinates the spindle assembly checkpoint (SAC), a signaling pathway that delays chromosome segregation until all sister chromatids are properly attached to spindle microtubules. Recently, microtubules and protein phosphatase 1 (PP1), which both bind the N-terminal domain of KNL1, have emerged as regulators of the SAC; however, how these proteins interact to contribute to SAC signaling is unknown. Here, we use X-ray crystallography, nuclear magnetic resonance spectroscopy, and biochemical assays to show how KNL1 binds both PP1 and microtubules. Unexpectedly, we discovered that PP1 and microtubules bind KNL1 via overlapping binding sites. Further, we showed that Aurora B kinase phosphorylation results in distinct patterns of KNL1 complex disruption. Finally, combining this data with co-sedimentation assays unequivocally demonstrated that microtubules and PP1 binding to KNL1 is mutually exclusive, with preferential formation of the KNL1:PP1 holoenzyme in the presence of PP1.
- Department of Chemistry and Biochemistry, University of Arizona, Tucson, AZ 85721, USA.
Organizational Affiliation: