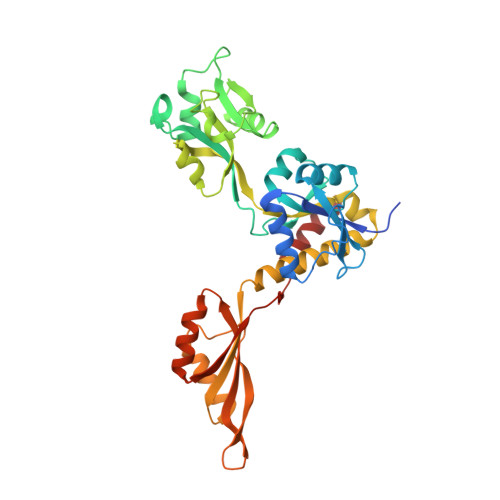
Guarding the gateway to histidine biosynthesis in plants:Medicago truncatulaATP-phosphoribosyltransferase in relaxed and tense states.
Ruszkowski, M.(2018) Biochem J 475: 2681-2697
- PubMed: 30072492 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20180289
- Primary Citation Related Structures:
6CZL, 6CZM - PubMed Abstract:
In the first committed step of histidine biosynthesis, adenosine 5'-triphosphate (ATP) and 5-phosphoribosyl-α1-pyrophosphate (PRPP), in the presence of ATP phosphoribosyltransferase (ATP-PRT, EC 2.4.2.17), yield phosphoribosyl-ATP. ATP-PRTs are subject to feedback inhibition by histidine that allosterically binds between the regulatory domains. Histidine biosynthetic pathways of bacteria, lower eukaryotes, and plants are considered promising targets for the design of antibiotics, antifungal agents, and herbicides because higher organisms are histidine heterotrophs. Plant ATP-PRTs are similar to one of the two types of their bacterial counterparts, the long-type ATP-PRTs. A biochemical and structural study of ATP-PRT from the model legume plant, Medicago truncatula ( Medtr ATP-PRT1) is reported herein. Two crystal structures, presenting homohexameric Medtr ATP-PRT1 in its relaxed (R-) and histidine-bound, tense (T-) states allowed to observe key features of the enzyme and provided the first structural insights into an ATP-PRT from a eukaryotic organism. In particular, they show pronounced conformational reorganizations during R-state to T-state transition that involves substantial movements of domains. This rearrangement requires a trans - to cis - switch of a peptide backbone within the hinge region of Medtr ATP-PRT1. A C-terminal α-helix, absent in bacteria, reinforces the hinge that is constituted by two peptide strands. As a result, conformations of the R- and T-states are significantly different from the corresponding states of prokaryotic enzymes with known 3-D structures. Finally, adenosine 5'-monophosphate (AMP) bound at the active site is consistent with a competitive (and synergistic with histidine) nature of AMP inhibition.
- Synchrotron Radiation Research Section of MCL, National Cancer Institute, S Cass Ave 9700, Argonne, IL, U.S.A. milosz.ruszkowski@nih.gov.
Organizational Affiliation: