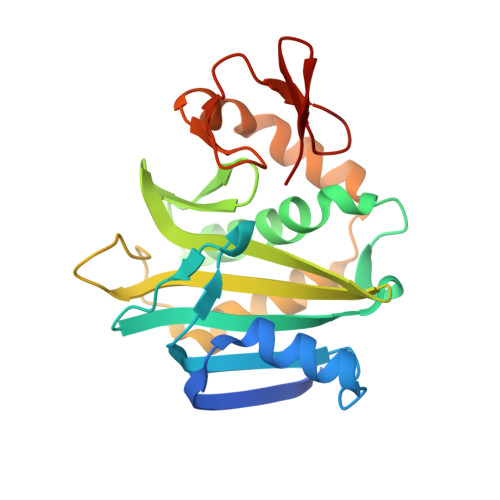
Co-crystal structure of Helicobacter pylori biotin protein ligase with biotinyl-5-ATP.
Ayanlade, J.P., Davis, D.E., Subramanian, S., Dranow, D.M., Lorimer, D.D., Hammerson, B., Myler, P.J., Asojo, O.A.(2025) Acta Crystallogr F Struct Biol Commun 81: 11-18
- PubMed: 39704719 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X24012056
- Primary Citation Related Structures:
6CK0 - PubMed Abstract:
Helicobacter pylori, a type 1 carcinogen that causes human gastric ulcers and cancer, is a priority target of the Seattle Structural Genomics Center for Infectious Disease (SSGCID). These efforts include determining the structures of potential H. pylori therapeutic targets. Here, the purification, crystallization and X-ray structure of one such target, H. pylori biotin protein ligase (HpBPL), are reported. HpBPL catalyzes the activation of various biotin-dependent metabolic pathways, including fatty-acid synthesis, gluconeogenesis and amino-acid catabolism, and may facilitate the survival of H. pylori in the high-pH gastric mucosa. HpBPL is a prototypical bacterial biotin protein ligase, despite having less than 35% sequence identity to any reported structure in the Protein Data Bank. A biotinyl-5-ATP molecule sits in a well conserved cavity. HpBPL shares extensive tertiary-structural similarity with Mycobacterium tuberculosis biotin protein ligase (MtBPL), despite having less than 22% sequence identity. The active site of HpBPL is very similar to that of MtBPL and has the necessary residues to bind inhibitors developed for MtBPL.
- Dartmouth Cancer Center, One Medical Center Drive, Lebanon, NH 03756, USA.
Organizational Affiliation: