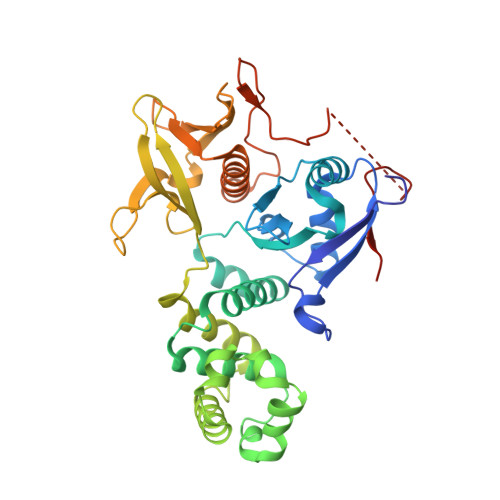
High resolution crystal structure of the FAK FERM domain reveals new insights on the Druggability of tyrosine 397 and the Src SH3 binding site.
Marlowe, T., Dementiev, A., Figel, S., Rivera, A., Flavin, M., Cance, W.(2019) BMC Mol Cell Biol 20: 10-10
- PubMed: 31109284 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s12860-019-0193-4
- Primary Citation Related Structures:
6CB0 - PubMed Abstract:
Focal Adhesion Kinase (FAK) is a major cancer drug target that is involved in numerous aspects of tumor progression and survival. While multiple research groups have developed ATP-competitive small molecule inhibitors that target the kinase enzyme, recent attention has been focused on the FAK FERM (Band 4.1, Ezrin, Radixin, Moesin) domain that contains key residue Y397 and contributes to many protein-protein interactions. Previous x-ray crystal structures of the FAK FERM domain gave conflicting results on the structure of the Y397 region and therefore the overall druggability. Here, we report the identification of a higher resolution crystal structure of the avian FAK FERM domain that shows conformational differences in Y397 and surrounding residues in the F1 lobe. In addition, we resolve the residues of the Src SH3 binding site, an area of the FERM domain that has previously shown limited electron density. These crystallographic data suggest that the Y397 region is highly dynamic and question the druggability of a putative pocket on the F1 lobe. In addition, new electron density data around the Src SH3 binding site provide structural insight on the FAK-Src activation cascade through a putative auto-inhibitory conformation.
- Interdisciplinary Oncology, University of Arizona College of Medicine - Phoenix, 475 N 5th Street, Phoenix, AZ, 85004, USA. tmarlowe@email.arizona.edu.
Organizational Affiliation: