Subtle Changes in the Combining Site of the Chlamydiaceae-Specific mAb S25-23 Increase the Antibody-Carbohydrate Binding Affinity by an Order of Magnitude.
Haji-Ghassemi, O., Muller-Loennies, S., Brooks, C.L., MacKenzie, C.R., Caveney, N., Van Petegem, F., Brade, L., Kosma, P., Brade, H., Evans, S.V.(2019) Biochemistry 58: 714-726
- PubMed: 30571096 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.8b00318
- Primary Citation Related Structures:
6C5H, 6C5I, 6C5J, 6C5K - PubMed Abstract:
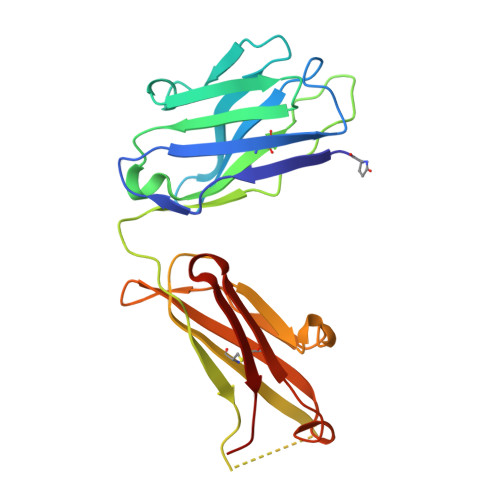
Murine antibodies S25-23, S25-26, and S25-5 derive from a common germ-line origin, and all bind the Chlamydiaceae family-specific epitope αKdo(2→8)αKdo(2→4)αKdo (where Kdo is 3-deoxy-α-d- manno-oct-2-ulosonic acid) with high affinity and specificity. These antibodies recognize the entire trisaccharide antigen in a linkage-dependent manner via a groove composed largely of germ-line residues. Despite sharing identical heavy and light chain genes, S25-23 binds the family-specific epitope with nanomolar affinity, which is an order of magnitude higher than that of S25-26, while S25-5 displays an affinity between those of S25-23 and S25-26. We determined the high-resolution crystal structures of S25-23 and S25-5 antigen binding fragments in complex with a pentasaccharide derived from the LPS of Chlamydia and measured the affinity of S25-5 for chlamydial LPS antigens using isothermal titration microcalorimetry. The 1.75 Å resolution structure of S25-23 shows how subtle conservative mutations Arg(L)-27E to lysine and Ser(H)-56 to threonine lead to an order of magnitude increase in affinity. Importantly, comparison between previous S25-26 structures and the 1.99 and 2.05 Å resolution liganded and unliganded structures of S25-5, respectively, shows how a Ser(L)-27E mutation results in an intermediate affinity due to the reduced enthalpic penalty associated with complex formation that would otherwise be required for arginine in this position. This strategy allows for subtle adjustments in the combining site via affinity maturation that have dramatic consequences for the affinity of an antibody for its antigen.
- Department of Biochemistry and Microbiology , University of Victoria , P.O. Box 3055 STN CSC, Victoria , British Columbia , Canada V8P 3P6.
Organizational Affiliation: