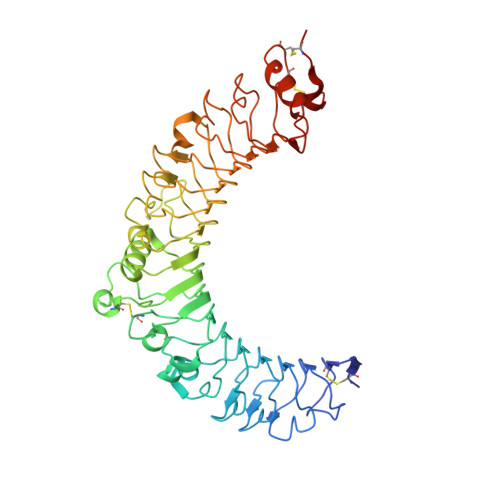
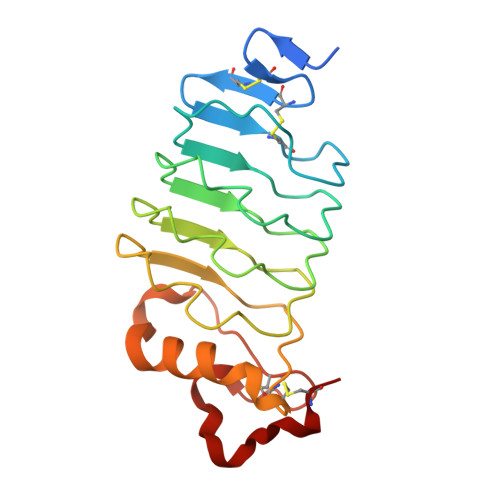
VLR Recognition of TLR5 Expands the Molecular Characterization of Protein Antigen Binding by Non-Ig-based Antibodies.
Gunn, R.J., Herrin, B.R., Acharya, S., Cooper, M.D., Wilson, I.A.(2018) J Mol Biology 430: 1350-1367
- PubMed: 29596914 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2018.03.016
- Primary Citation Related Structures:
6BXA, 6BXC, 6BXD, 6BXE - PubMed Abstract:
Variable lymphocyte receptors (VLRs) are unconventional adaptive immune receptors relatively recently discovered in the phylogenetically ancient jawless vertebrates, lamprey and hagfish. VLRs bind antigens using a leucine-rich repeat fold and are the only known adaptive immune receptors that do not utilize an immunoglobulin fold for antigen recognition. While immunoglobulin antibodies have been studied extensively, there are comparatively few studies on antigen recognition by VLRs, particularly for protein antigens. Here we report isolation, functional and structural characterization of three VLRs that bind the protein toll-like receptor 5 (TLR5) from zebrafish. Two of the VLRs block binding of TLR5 to its cognate ligand flagellin in functional assays using reporter cells. Co-crystal structures revealed that these VLRs bind to two different epitopes on TLR5, both of which include regions involved in flagellin binding. Our work here demonstrates that the lamprey adaptive immune system can be used to generate high-affinity VLR clones that recognize different epitopes and differentially impact natural ligand binding to a protein antigen.
- Department of Integrative Structural and Computational Biology and the Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: