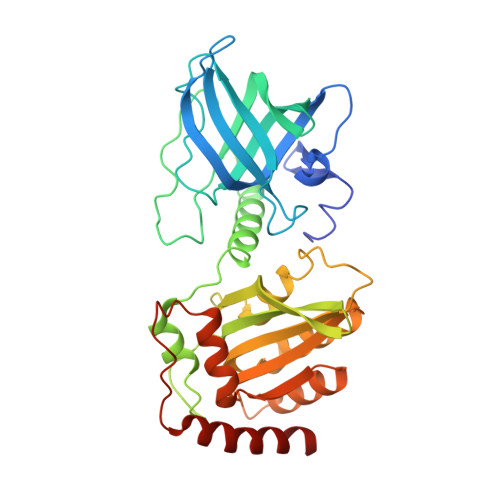
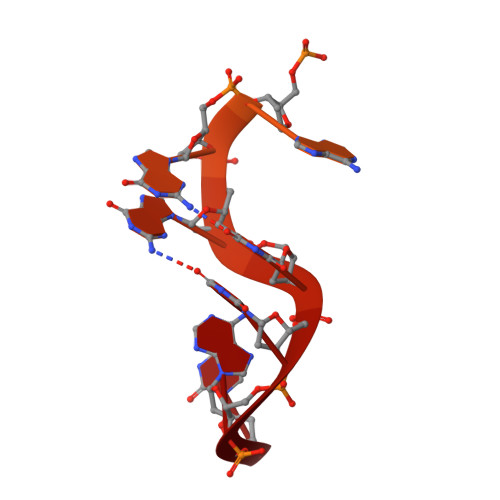
Insights into DNA substrate selection by APOBEC3G from structural, biochemical, and functional studies.
Ziegler, S.J., Liu, C., Landau, M., Buzovetsky, O., Desimmie, B.A., Zhao, Q., Sasaki, T., Burdick, R.C., Pathak, V.K., Anderson, K.S., Xiong, Y.(2018) PLoS One 13: e0195048-e0195048
- PubMed: 29596531 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0195048
- Primary Citation Related Structures:
6BWY - PubMed Abstract:
Human apolipoprotein B mRNA-editing enzyme-catalytic polypeptide-like 3 (A3) proteins are a family of cytidine deaminases that catalyze the conversion of deoxycytidine (dC) to deoxyuridine (dU) in single-stranded DNA (ssDNA). A3 proteins act in the innate immune response to viral infection by mutating the viral ssDNA. One of the most well-studied human A3 family members is A3G, which is a potent inhibitor of HIV-1. Each A3 protein prefers a specific substrate sequence for catalysis-for example, A3G deaminates the third dC in the CCCA sequence motif. However, the interaction between A3G and ssDNA is difficult to characterize due to poor solution behavior of the full-length protein and loss of DNA affinity of the truncated protein. Here, we present a novel DNA-anchoring fusion strategy using the protection of telomeres protein 1 (Pot1) which has nanomolar affinity for ssDNA, with which we captured an A3G-ssDNA interaction. We crystallized a non-preferred adenine in the -1 nucleotide-binding pocket of A3G. The structure reveals a unique conformation of the catalytic site loops that sheds light onto how the enzyme scans substrate in the -1 pocket. Furthermore, our biochemistry and virology studies provide evidence that the nucleotide-binding pockets on A3G influence each other in selecting the preferred DNA substrate. Together, the results provide insights into the mechanism by which A3G selects and deaminates its preferred substrates and help define how A3 proteins are tailored to recognize specific DNA sequences. This knowledge contributes to a better understanding of the mechanism of DNA substrate selection by A3G, as well as A3G antiviral activity against HIV-1.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut, United States of America.
Organizational Affiliation: