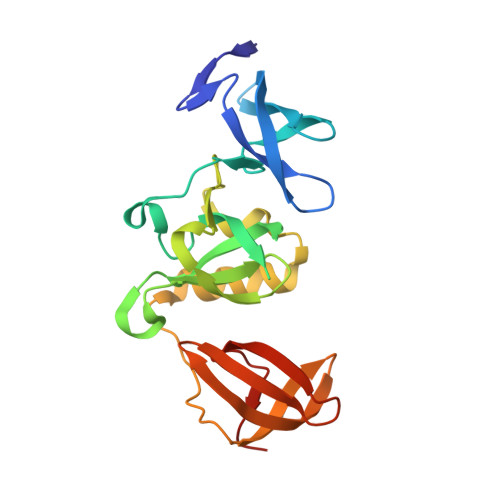
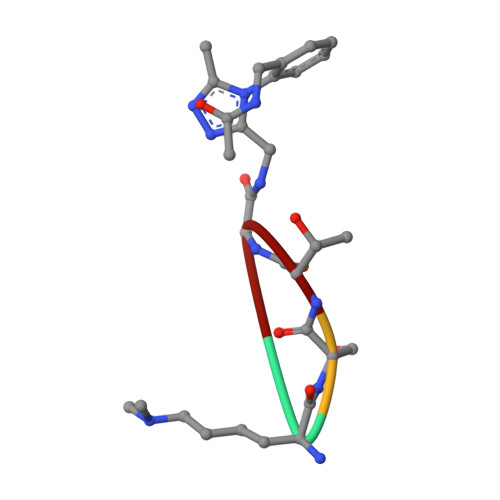
Crystal structure of SETDB1 Tudor domain with aryl triazole fragment peptide conjugates
MADER, P., Mendoza-Sanchez, R., DONG, A., DOBROVETSKY, E., IQBAL, A., CORLESS, V., TEMPEL, W., LIEW, S.K., SMIL, D., DELA SENA, C.C., KENNEDY, S., DIAZ, D.B., SCHAPIRA, M., VEDADI, M., BROWN, P.J., Santhakumar, V., FRYE, S., Bountra, C., Edwards, A.M., YUDIN, A.K., Arrowsmith, C.H., Structural Genomics Consortium (SGC)To be published.