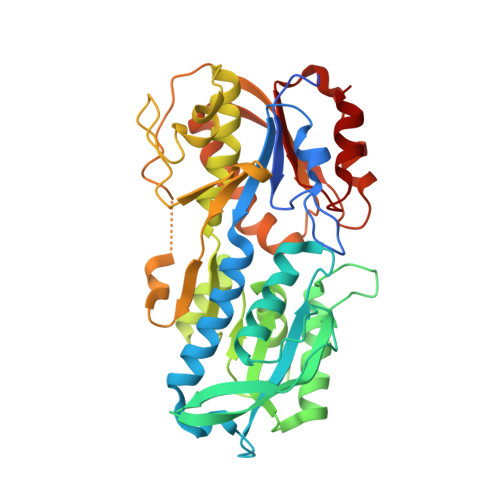
Crystal structures of MglB-2 (TP0684), a topologically variant d-glucose-binding protein from Treponema pallidum, reveal a ligand-induced conformational change.
Brautigam, C.A., Deka, R.K., Liu, W.Z., Norgard, M.V.(2018) Protein Sci 27: 880-885
- PubMed: 29318719 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3373
- Primary Citation Related Structures:
6BGC, 6BGD - PubMed Abstract:
Previously, we determined the crystal structure of apo-TpMglB-2, a d-glucose-binding component of a putative ABC transporter from the syphilis spirochete Treponema pallidum. The protein had an unusual topology for this class of proteins, raising the question of whether the d-glucose-binding mode would be different in TpMglB-2. Here, we present the crystal structures of a variant of TpMglB-2 with and without d-glucose bound. The structures demonstrate that, despite its aberrant topology, the protein undergoes conformational changes and binds d-glucose similarly to other Mgl-type proteins, likely facilitating d-glucose uptake in T. pallidum.
- Department of Biophysics, The University of Texas Southwestern Medical Center, Dallas, Texas, 75390.
Organizational Affiliation: