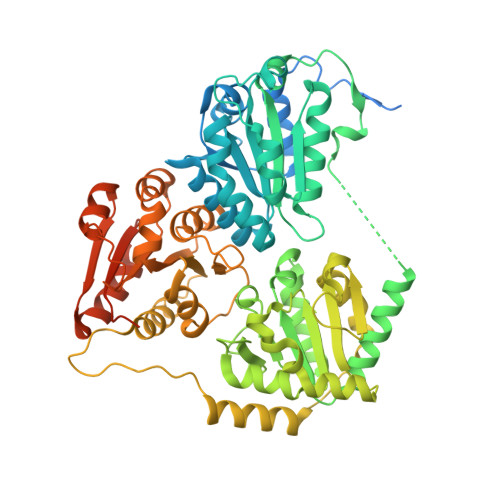
High resolution crystal structures of the acetohydroxyacid synthase-pyruvate complex provide new insights into its catalytic mechanism
Lonhienne, T., Garcia, M.D., Noble, C., Harmer, J., Fraser, J.A., Williams, C.M., Guddat, L.W.(2017) ChemistrySelect