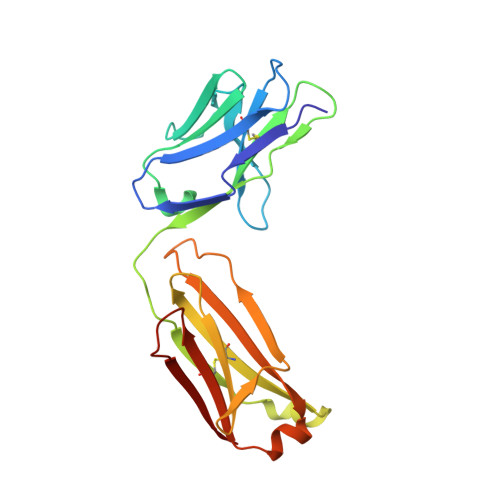
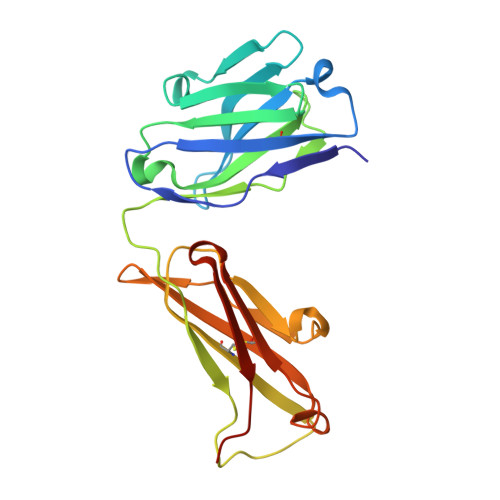
Tau Antibody Structure Reveals a Molecular Switch Defining a Pathological Conformation of the Tau Protein.
Chukwu, J.E., Pedersen, J.T., Pedersen, L.O., Volbracht, C., Sigurdsson, E.M., Kong, X.P.(2018) Sci Rep 8: 6209-6209
- PubMed: 29670132 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-24276-4
- Primary Citation Related Structures:
6BB4 - PubMed Abstract:
Tau antibodies have shown therapeutic potential for Alzheimer's disease and several are in clinical trials. As a microtubule-associated protein, tau relies on dynamic phosphorylation for its normal functions. In tauopathies, it becomes hyperphosphorylated and aggregates into toxic assemblies, which collectively lead to neurodegeneration. Of the phospho-epitopes, the region around Ser396 has received particular attention because of its prominence and stability in tauopathies. Here we report the first structure of a monoclonal tau antibody in complex with the pathologically important phospho-Ser396 residue. Its binding region reveals tau residues Tyr394 to phospho-Ser396 stabilized in a β-strand conformation that is coordinated by a phospho-specific antigen binding site. These details highlight a molecular switch that defines this prominent conformation of tau and ways to target it. Overall, the structure of the antibody-antigen complex clarifies why certain phosphorylation sites in tau are more closely linked to neurodegeneration than others.
- Departments of Biochemistry & Molecular Pharmacology, New York University School of Medicine, New York, NY, USA.
Organizational Affiliation: