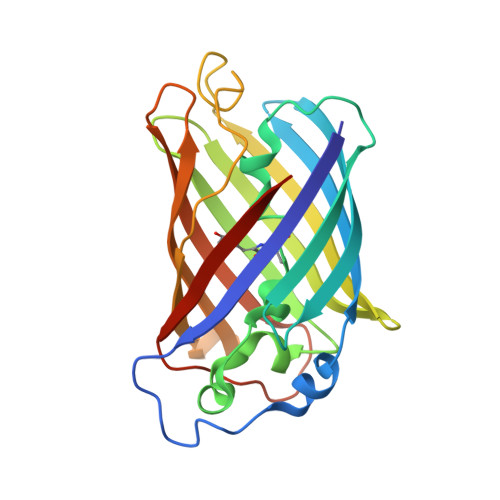
Structural Insight into the Photochemistry of Split Green Fluorescent Proteins: A Unique Role for a His-Tag.
Deng, A., Boxer, S.G.(2018) J Am Chem Soc 140: 375-381
- PubMed: 29193968 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.7b10680
- Primary Citation Related Structures:
6B7R, 6B7T - PubMed Abstract:
Oligohistidine affinity tags (His-tags) are commonly fused to proteins to aid in their purification via metal affinity chromatography. These His-tags are generally assumed to have minimal impact on the properties of the fusion protein, as they have no propensity to form ordered elements, and are small enough not to significantly affect the solubility or size. Here we report structures of two variants of truncated green fluorescent protein (GFP), i.e., split GFP with a β-strand removed, that were found to behave differently in the presence of light. In these structures, the N-terminal His-tag and several neighboring residues play a highly unusual structural and functional role in stabilizing the truncated GFP by substituting as a surrogate β-strand in the groove vacated by the native strand. This finding provides an explanation for the seemingly very different peptide binding and photodissociation properties of split proteins involving β-strands 10 and 11. We show that these truncated GFPs can bind other non-native sequences, and this promiscuity invites the possibility for rational design of sequences optimized for strand binding and photodissociation, both useful for optogenetic applications.
- Department of Chemistry, Stanford University , Stanford, California 94305-5012, United States.
Organizational Affiliation: