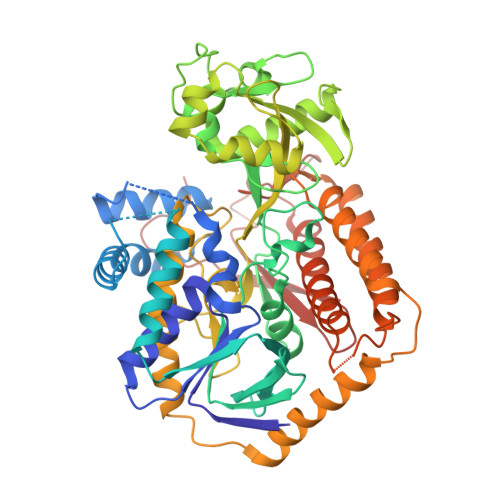
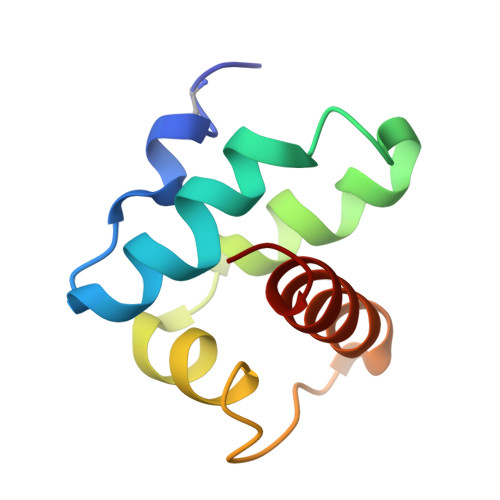
Crystal structure of an assembly intermediate of respiratory Complex II.
Sharma, P., Maklashina, E., Cecchini, G., Iverson, T.M.(2018) Nat Commun 9: 274-274
- PubMed: 29348404 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02713-8
- Primary Citation Related Structures:
6B58 - PubMed Abstract:
Flavin is covalently attached to the protein scaffold in ~10% of flavoenzymes. However, the mechanism of covalent modification is unclear, due in part to challenges in stabilizing assembly intermediates. Here, we capture the structure of an assembly intermediate of the Escherichia coli Complex II (quinol:fumarate reductase (FrdABCD)). The structure contains the E. coli FrdA subunit bound to covalent FAD and crosslinked with its assembly factor, SdhE. The structure contains two global conformational changes as compared to prior structures of the mature protein: the rotation of a domain within the FrdA subunit, and the destabilization of two large loops of the FrdA subunit, which may create a tunnel to the active site. We infer a mechanism for covalent flavinylation. As supported by spectroscopic and kinetic analyses, we suggest that SdhE shifts the conformational equilibrium of the FrdA active site to disfavor succinate/fumarate interconversion and enhance covalent flavinylation.
- Department of Pharmacology, Vanderbilt University, Nashville, TN, 37232, USA.
Organizational Affiliation: