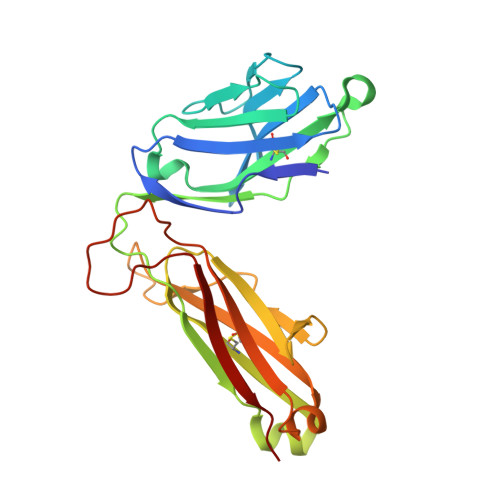
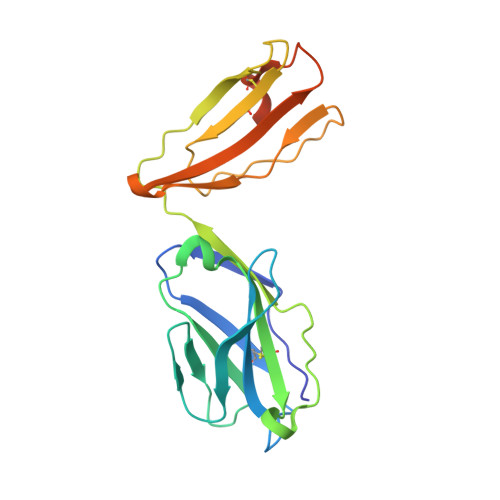
Divergent T-cell receptor recognition modes of a HLA-I restricted extended tumour-associated peptide.
Chan, K.F., Gully, B.S., Gras, S., Beringer, D.X., Kjer-Nielsen, L., Cebon, J., McCluskey, J., Chen, W., Rossjohn, J.(2018) Nat Commun 9: 1026-1026
- PubMed: 29531227 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-03321-w
- Primary Citation Related Structures:
6AT5, 6AT6, 6AVF, 6AVG - PubMed Abstract:
Human leukocyte antigen (HLA)-I molecules generally bind short peptides (8-10 amino acids), although extended HLA-I restricted peptides (>10 amino acids) can be presented to T cells. However, the function of such extended HLA-I epitopes in tumour immunity, and how they would be recognised by T-cell receptors (TCR) remains unclear. Here we show that the structures of two distinct TCRs (TRAV4 + TRAJ21 + -TRBV28 + TRBJ2-3 + and TRAV4 + TRAJ8 + -TRBV9 + TRBJ2-1 + ), originating from a polyclonal T-cell repertoire, bind to HLA-B*07:02, presenting a 13-amino-acid-long tumour-associated peptide, NY-ESO-1 60-72 . Comparison of the structures reveals that the two TCRs differentially binds NY-ESO-1 60-72 -HLA-B*07:02 complex, and induces differing extent of conformational change of the NY-ESO-1 60-72 epitope. Accordingly, polyclonal TCR usage towards an extended HLA-I restricted tumour epitope translates to differing TCR recognition modes, whereby extensive flexibility at the TCR-pHLA-I interface engenders recognition.
- Olivia Newton-John Cancer Research Institute, and School of Cancer Medicine, La Trobe University, Heidelberg, VIC, 3084, Australia.
Organizational Affiliation: