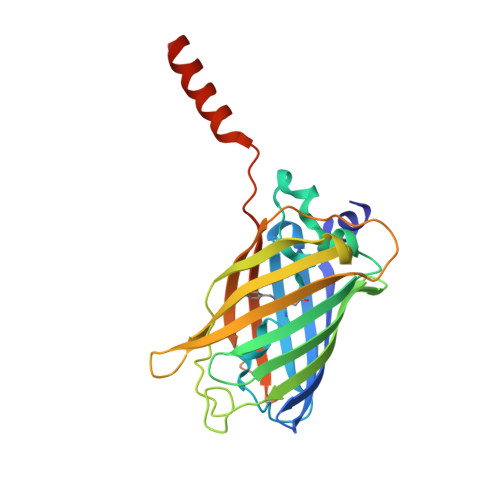
Atomic insights into the genesis of cellular filaments by globular proteins.
McPartland, L., Heller, D.M., Eisenberg, D.S., Hochschild, A., Sawaya, M.R.(2018) Nat Struct Mol Biol 25: 705-714
- PubMed: 30076408 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-018-0096-7
- Primary Citation Related Structures:
6AS9 - PubMed Abstract:
Self-assembly of proteins into filaments, such as actin and tubulin filaments, underlies essential cellular processes in all three domains of life. The early emergence of filaments in evolutionary history suggests that filament genesis might be a robust process. Here we describe the fortuitous construction of GFP fusion proteins that self-assemble as fluorescent polar filaments in Escherichia coli. Filament formation is achieved by appending as few as 12 residues to GFP. Crystal structures reveal that each protomer donates an appendage to fill a groove between the two following protomers along the filament. This exchange of appendages resembles runaway domain swapping but is distinguished by higher efficiency because monomers cannot competitively bind their own appendages. Ample evidence for this 'runaway domain coupling' mechanism in nature suggests it could facilitate the evolutionary pathway from globular protein to polar filament, requiring a minimal extension of protein sequence and no substantial refolding.
- Department of Microbiology and Immunobiology, Harvard Medical School, Boston, MA, USA.
Organizational Affiliation: