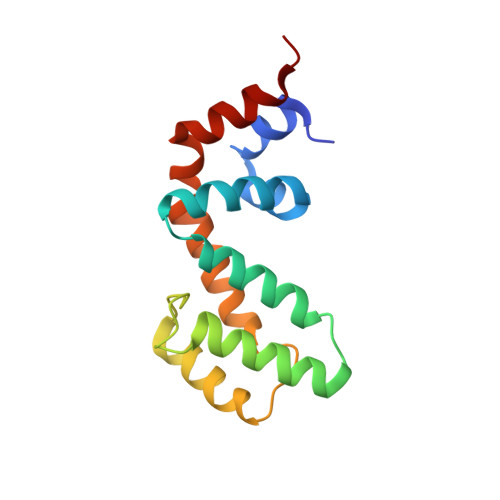
High-resolution structure of RGS17 suggests a role for Ca2+in promoting the GTPase-activating protein activity by RZ subfamily members.
Sieng, M., Hayes, M.P., O'Brien, J.B., Andrew Fowler, C., Houtman, J.C., Roman, D.L., Lyon, A.M.(2019) J Biological Chem 294: 8148-8160
- PubMed: 30940727 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.006059
- Primary Citation Related Structures:
6AM3 - PubMed Abstract:
Regulator of G protein signaling (RGS) proteins are negative regulators of G protein-coupled receptor (GPCR) signaling through their ability to act as GTPase-activating proteins (GAPs) for activated Gα subunits. Members of the RZ subfamily of RGS proteins bind to activated Gα o , Gα z , and Gα i1-3 proteins in the nervous system and thereby inhibit downstream pathways, including those involved in Ca 2+ -dependent signaling. In contrast to other RGS proteins, little is known about RZ subfamily structure and regulation. Herein, we present the 1.5-Å crystal structure of RGS17, the most complete and highest-resolution structure of an RZ subfamily member to date. RGS17 cocrystallized with Ca 2+ bound to conserved positions on the predicted Gα-binding surface of the protein. Using NMR chemical shift perturbations, we confirmed that Ca 2+ binds in solution to the same site. Furthermore, RGS17 had greater than 55-fold higher affinity for Ca 2+ than for Mg 2+ Finally, we found that Ca 2+ promotes interactions between RGS17 and activated Gα and decreases the K m for GTP hydrolysis, potentially by altering the binding mechanism between these proteins. Taken together, these findings suggest that Ca 2+ positively regulates RGS17, which may represent a general mechanism by which increased Ca 2+ concentration promotes the GAP activity of the RZ subfamily, leading to RZ-mediated inhibition of Ca 2+ signaling.
- Department of Chemistry, Purdue University, West Lafayette, Indiana 47907.
Organizational Affiliation: