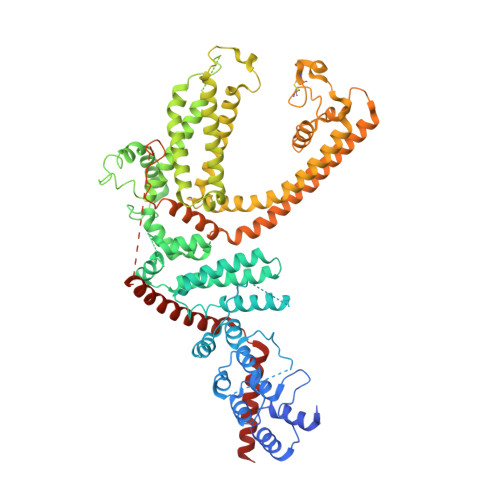
Cryo-EM structure of TRPC5 at 2.8- angstrom resolution reveals unique and conserved structural elements essential for channel function.
Duan, J., Li, J., Chen, G.L., Ge, Y., Liu, J., Xie, K., Peng, X., Zhou, W., Zhong, J., Zhang, Y., Xu, J., Xue, C., Liang, B., Zhu, L., Liu, W., Zhang, C., Tian, X.L., Wang, J., Clapham, D.E., Zeng, B., Li, Z., Zhang, J.(2019) Sci Adv 5
- PubMed: 31355338 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aaw7935
- Primary Citation Related Structures:
6AEI - PubMed Abstract:
The transient receptor potential canonical subfamily member 5 (TRPC5), one of seven mammalian TRPC members, is a nonselective calcium-permeant cation channel. TRPC5 is of considerable interest as a drug target in the treatment of progressive kidney disease, depression, and anxiety. Here, we present the 2.8-Å resolution cryo-electron microscopy (cryo-EM) structure of the mouse TRPC5 (mTRPC5) homotetramer. Comparison of the TRPC5 structure to previously determined structures of other TRPC and TRP channels reveals differences in the extracellular pore domain and in the length of the S3 helix. The disulfide bond at the extracellular side of the pore and a preceding small loop are essential elements for its proper function. This high-resolution structure of mTRPC5, combined with electrophysiology and mutagenesis, provides insight into the lipid modulation and gating mechanisms of the TRPC family of ion channels.
- Human Aging Research Institute (HARI), School of Life Sciences, Nanchang University, Nanchang, Jiangxi 330031, China.
Organizational Affiliation: