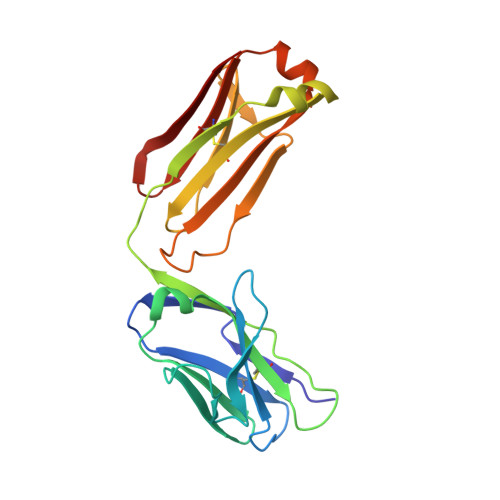
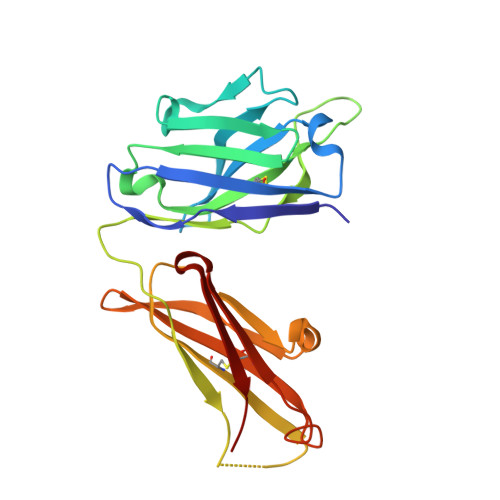
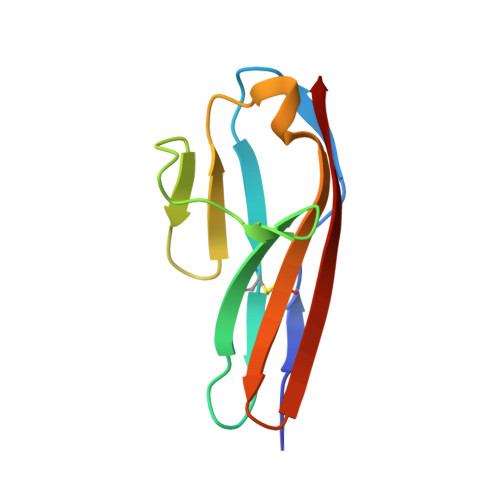Affinity Improvement of a Cancer-Targeted Antibody through Alanine-Induced Adjustment of Antigen-Antibody Interface.
Yamashita, T., Mizohata, E., Nagatoishi, S., Watanabe, T., Nakakido, M., Iwanari, H., Mochizuki, Y., Nakayama, T., Kado, Y., Yokota, Y., Matsumura, H., Kawamura, T., Kodama, T., Hamakubo, T., Inoue, T., Fujitani, H., Tsumoto, K.(2019) Structure 27: 519
- PubMed: 30595454 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2018.11.002
- Primary Citation Related Structures:
6A76, 6A77, 6A78, 6A79 - PubMed Abstract:
To investigate favorable single amino acid substitutions that improve antigen-antibody interactions, alanine (Ala) mutagenesis scanning of the interfacial residues of a cancer-targeted antibody, B5209B, was performed based on X-ray crystallography analysis. Two substitutions were shown to significantly enhance the binding affinity for the antigen, by up to 30-fold. One substitution improved the affinity by a gain of binding enthalpy, whereas the other substitution improved the affinity by a gain of binding entropy. Molecular dynamics simulations showed that the enthalpic improvement could be attributed to the stabilization of distant salt bridges located at the periphery of the antigen-antibody interface. The entropic improvement was due to the release of water molecules that were stably trapped in the antigen-antibody interface of the wild-type antibody. Importantly, these effects of the Ala substitutions were caused by subtle adjustments of the binding interface. These results will be helpful to design high-affinity antibodies with avoiding entropy-enthalpy compensation.
- Laboratory for Systems Biology and Medicine, Research Center for Advanced Science and Technology, The University of Tokyo, 4-6-1, Komaba, Meguro-ku, Tokyo 153-8904, Japan.
Organizational Affiliation: