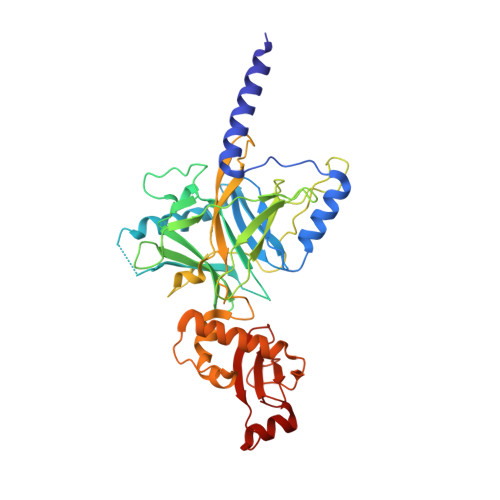
Crystal structure of the central and the C-terminal RNase domains of colicin D implicated its translocation pathway through inner membrane of target cell
Chang, J.W., Sato, Y., Ogawa, T., Arakawa, T., Fukai, S., Fushinobu, S., Masaki, H.(2018) J Biochem 164: 329-339
- PubMed: 29905832 Search on PubMed
- DOI: https://doi.org/10.1093/jb/mvy056
- Primary Citation Related Structures:
5ZNM - PubMed Abstract:
Colicins are protein toxins produced by and toxic to Escherichia coli strains. Colicin D consists of an N-terminal domain (NTD), central domain (CD) and C-terminal RNase domain (CRD). The cognate immunity protein, ImmD, is co-synthesized in producer cells to block the toxic tRNase activity of the CRD. Previous studies have reported the crystal structure of CRD/ImmD complex. Colicin D hijacks the surface receptor FepA and the energy transducer TonB system using the NTD for translocation across the outer membrane of the target cells. The CD is required for endoproteolytic processing and the translocation of CRD across the inner membrane, and the membrane-associated protease FtsH and the signal peptidase LepB are exploited in this process. Although several regions of the CD have been identified in interactions with the hijacked inner membrane system or immunity protein, the structural basis of the CD is unknown. In this study, we determined the crystal structure of colicin D, containing both the CD and CRD. The full-length colicin D/ImmD heterodimer structure was built by superimposing the CD-CRD structure with the previously determined partial structures. The overall translocation process of colicin D, including the interaction between CD and LepB, is discussed.
- Department of Biotechnology, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo, Japan.
Organizational Affiliation: