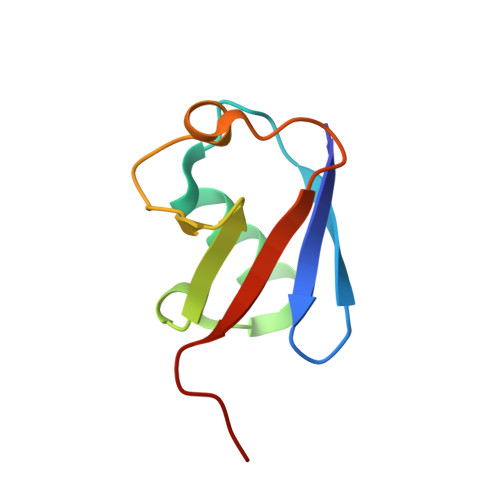
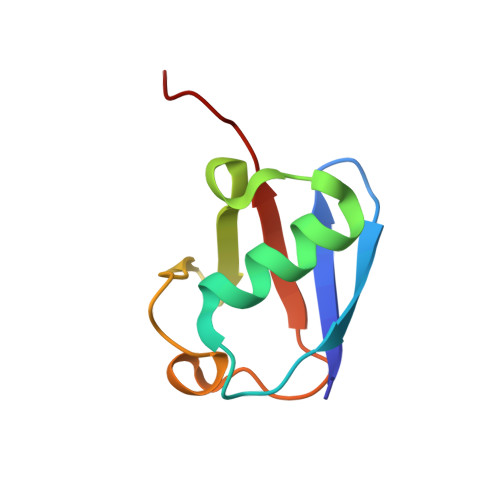
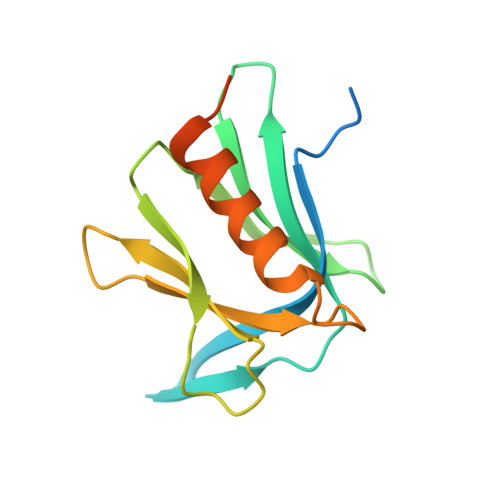
Structural basis for the recognition of K48-linked Ub chain by proteasomal receptor Rpn13.
Liu, Z., Dong, X., Yi, H.W., Yang, J., Gong, Z., Wang, Y., Liu, K., Zhang, W.P., Tang, C.(2019) Cell Discov 5: 19-19
- PubMed: 30962947 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-019-0089-7
- Primary Citation Related Structures:
5YMY - PubMed Abstract:
The interaction between K48-linked ubiquitin (Ub) chain and Rpn13 is important for proteasomal degradation of ubiquitinated substrate proteins. Only the complex structure between the N-terminal domain of Rpn13 (Rpn13 NTD ) and Ub monomer has been characterized, while it remains unclear how Rpn13 specifically recognizes K48-linked Ub chain. Using single-molecule FRET, here we show that K48-linked diubiquitin (K48-diUb) fluctuates among distinct conformational states, and a preexisting compact state is selectively enriched by Rpn13 NTD . The same binding mode is observed for full-length Rpn13 and longer K48-linked Ub chain. Using solution NMR spectroscopy, we have determined the complex structure between Rpn13 NTD and K48-diUb. In this structure, Rpn13 NTD simultaneously interacts with proximal and distal Ub subunits of K48-diUb that remain associated in the complex, thus corroborating smFRET findings. The proximal Ub interacts with Rpn13 NTD similarly as the Ub monomer in the known Rpn13 NTD :Ub structure, while the distal Ub binds to a largely electrostatic surface of Rpn13 NTD . Thus, a charge-reversal mutation in Rpn13 NTD weakens the interaction between Rpn13 and K48-linked Ub chain, causing accumulation of ubiquitinated proteins. Moreover, physical blockage of the access of the distal Ub to Rpn13 NTD with a proximity-attached Ub monomer can disrupt the interaction between Rpn13 and K48-diUb. Taken together, the bivalent interaction of K48-linked Ub chain with Rpn13 provides the structural basis for Rpn13 linkage selectivity, which opens a new window for modulating proteasomal function.
- 1CAS Key Laboratory of Magnetic Resonance in Biological Systems, State Key Laboratory of Magnetic Resonance and Atomic Molecular Physics, National Center for Magnetic Resonance at Wuhan, Wuhan Institute of Physics and Mathematics of the Chinese Academy of Sciences, Wuhan, Hubei Province 430071 China.
Organizational Affiliation: